interpElections provides 16 diagnostic functions for validating interpolation results. This vignette demonstrates each one using real data from four Brazilian municipalities of varying sizes:
| City | State | Tracts | Sources | Character |
|---|---|---|---|---|
| Igrejinha | RS | ~40 | ~10 | Tiny rural municipality |
| Varginha | MG | ~280 | ~37 | Small city (primary example) |
| Niteroi | RJ | ~450 | ~80 | Medium, dense urban area |
| Belo Horizonte | MG | ~5,100 | ~600 | Major metropolitan area |
Each section follows the pattern: Intuition (why it matters), Formal definition (the math), Applied example (real output), Interpretation (what to look for).
Setup
library(interpElections)
# Load precomputed results (2022 presidential election)
result_igr <- readRDS("precomputed/igrejinha_2022.rds")
result_vga <- readRDS("precomputed/varginha_2022.rds")
result_vga2 <- readRDS("precomputed/varginha_2022_t2.rds") # turno 2
result_nit <- readRDS("precomputed/niteroi_2022.rds")
result_bh <- readRDS("precomputed/belo_horizonte_2022.rds")Quick overview of each result:
#> interpElections result summary
#> --------------------------------------------------
#> Municipality: VARGINHA (MG) -- IBGE: 3170701, TSE: 54135
#> Election: 2022 | Census: 2022 | Turno: 1 | Cargo: PRESIDENTE
#> Census tracts: 279 | Sources: 37 | Variables: 28
#>
#> Calibration: gender x 7 age brackets (14 pairs)
#>
#> Optimization: pb_sgd_colnorm_cpu | Objective: 4670.4370 | Convergence: 0
#> Alpha: min=1.000, Q1=1.604, median=1.870, Q3=2.227, max=11.836
#>
#> Candidates (13):
#> CAND_30 LUIZ FELIPE CHAVES D AVILA (NOVO) total= NA mean= NA [NA, NA]
#> CAND_44 SORAYA VIEIRA THRONICKE (UNIÃO) total= NA mean= NA [NA, NA]
#> CAND_12 CIRO FERREIRA GOMES (PDT) total= NA mean= NA [NA, NA]
#> CAND_13 LUIZ INÁCIO LULA DA SILVA (PT) total= NA mean= NA [NA, NA]
#> ... 9 more -- View(result$dictionary)
#>
#> Turnout (1):
#> QT_COMPARECIMENTO turnout total= NA mean= NA [NA, NA]
#>
#> Calibration (14):
#> vot_hom_18_19 calibration total= NA mean= NA [NA, NA]
#> vot_hom_20_24 calibration total= NA mean= NA [NA, NA]
#> vot_hom_25_29 calibration total= NA mean= NA [NA, NA]
#> vot_hom_30_39 calibration total= NA mean= NA [NA, NA]
#> vot_hom_40_49 calibration total= NA mean= NA [NA, NA]
#> vot_hom_50_59 calibration total= NA mean= NA [NA, NA]
#> vot_hom_60_69 calibration total= NA mean= NA [NA, NA]
#> vot_mul_18_19 calibration total= NA mean= NA [NA, NA]
#> vot_mul_20_24 calibration total= NA mean= NA [NA, NA]
#> vot_mul_25_29 calibration total= NA mean= NA [NA, NA]
#> vot_mul_30_39 calibration total= NA mean= NA [NA, NA]
#> vot_mul_40_49 calibration total= NA mean= NA [NA, NA]
#> vot_mul_50_59 calibration total= NA mean= NA [NA, NA]
#> vot_mul_60_69 calibration total= NA mean= NA [NA, NA]
#>
#>
#> Object size: 181.3 MB
#> Neighborhoods: 387#> interpElections result summary
#> --------------------------------------------------
#> Municipality: BELO HORIZONTE (MG) -- IBGE: 3106200, TSE: 41238
#> Election: 2022 | Census: 2022 | Turno: 1 | Cargo: PRESIDENTE
#> Census tracts: 5109 | Sources: 407 | Variables: 28
#>
#> Calibration: gender x 7 age brackets (14 pairs)
#>
#> Optimization: pb_sgd_colnorm_cuda | Objective: 13281.0293 | Convergence: 0
#> Alpha: min=1.000, Q1=1.248, median=1.357, Q3=1.487, max=17.115
#>
#> Candidates (13):
#> CAND_13 LUIZ INÁCIO LULA DA SILVA (PT) total= 603542 mean= 118.1 [2.8, 334.0]
#> CAND_27 JOSE MARIA EYMAEL (DC) total= 209 mean= 0.0 [0.0, 0.3]
#> CAND_30 LUIZ FELIPE CHAVES D AVILA (NOVO) total= 14496 mean= 2.8 [0.1, 12.2]
#> CAND_14 KELMON LUIS DA SILVA SOUZA (PTB) total= 861 mean= 0.2 [0.0, 1.0]
#> ... 9 more -- View(result$dictionary)
#>
#> Turnout (1):
#> QT_COMPARECIMENTO turnout total= 1489578 mean= 291.6 [7.1, 802.2]
#>
#> Calibration (14):
#> vot_hom_18_19 calibration total= 21044 mean= 4.1 [0.0, 12.6]
#> vot_hom_20_24 calibration total= 73593 mean= 14.4 [0.3, 44.4]
#> vot_hom_25_29 calibration total= 81481 mean= 15.9 [0.4, 58.2]
#> vot_hom_30_39 calibration total= 172752 mean= 33.8 [0.8, 102.2]
#> vot_hom_40_49 calibration total= 170766 mean= 33.4 [0.8, 113.1]
#> vot_hom_50_59 calibration total= 138388 mean= 27.1 [0.7, 71.7]
#> vot_hom_60_69 calibration total= 105049 mean= 20.6 [0.4, 52.7]
#> vot_mul_18_19 calibration total= 23556 mean= 4.6 [0.0, 15.0]
#> vot_mul_20_24 calibration total= 77657 mean= 15.2 [0.3, 46.7]
#> vot_mul_25_29 calibration total= 88270 mean= 17.3 [0.4, 69.7]
#> vot_mul_30_39 calibration total= 190378 mean= 37.3 [0.9, 110.6]
#> vot_mul_40_49 calibration total= 194436 mean= 38.1 [0.9, 136.5]
#> vot_mul_50_59 calibration total= 165447 mean= 32.4 [0.8, 86.6]
#> vot_mul_60_69 calibration total= 137434 mean= 26.9 [0.6, 68.5]
#>
#>
#> Object size: 243.8 MB
#> Neighborhoods: 8651. Automated Health Check: diagnostics()
Intuition
diagnostics() is a single-call “blood test” for your
interpolation result. It runs 8 automated checks and reports
PASS/WARN/FAIL for each. Use it as a first step after every
interpolation to catch problems early.
Only checks whose outcome is not guaranteed by construction are included. Vote conservation, non-negativity, and column normalization are all enforced by design and therefore omitted.
Formal definition
| Check | Statistic | PASS | WARN | FAIL |
|---|---|---|---|---|
| Convergence | Optimizer exit code | code = 0 | code != 0 | – |
| Alpha distribution | % of tracts with | < 5% | >= 5% | – |
| Residual RMSE | always (informational) | – | – | |
| Residual bias | ratio < 10% | ratio >= 10% | – | |
| Pop-voter gap | < 5% | 5–20% | > 20% | |
| Unreachable pairs | % of near max | < 1% | 1–10% | > 10% |
| Alpha spatial CV | CV >= 0.1 | CV < 0.1 | – | |
| Implied turnout | % of tracts with rate > 110% | < 5% | >= 5% | – |
Applied example
diagnostics(result_vga)#> interpElections diagnostics
#> --------------------------------------------------
#> [PASS] Convergence: optimizer converged (code 0)
#> [PASS] Alpha distribution: median=1.83, IQR=[1.61, 2.16], 0% > 15
#> [PASS] Residual RMSE: 11.3331 (informational)
#> [WARN] Residual bias: max |colMeans(resid)| = 8.6344 (76.2% of RMSE)
#> [WARN] Population-voter gap: census=96829, source=78389, gap=19.0%
#> [WARN] Unreachable pairs: 2.9% of pairs unreachable (298 / 10323 NA)
#> [PASS] Alpha spatial variation: CV of median-alpha = 0.492
#> [PASS] Implied turnout: 1.1% of tracts with implied turnout > 110%
#> --------------------------------------------------
#> 5 PASS, 3 WARNFor a larger city:
diagnostics(result_bh)#> interpElections diagnostics
#> --------------------------------------------------
#> [PASS] Convergence: optimizer converged (code 0)
#> [PASS] Alpha distribution: median=1.36, IQR=[1.26, 1.47], 0% > 15
#> [PASS] Residual RMSE: 7.2604 (informational)
#> [WARN] Residual bias: max |colMeans(resid)| = 1.8894 (26.0% of RMSE)
#> [PASS] Population-voter gap: census=1650864, source=1640251, gap=0.6%
#> [FAIL] Unreachable pairs: 48.0% of pairs unreachable (997611 / 2079363 NA)
#> [PASS] Alpha spatial variation: CV of median-alpha = 0.567
#> [PASS] Implied turnout: 2.2% of tracts with implied turnout > 110%
#> --------------------------------------------------
#> 6 PASS, 1 WARN, 1 FAILYou can also access check values programmatically:
d <- diagnostics(result_vga, verbose = FALSE)
d$checks$convergence$value
d$checks$residual_rmse$value
d$checks$pop_voter_gap$valueInterpretation
-
Convergence WARN: The optimizer didn’t converge
within the epoch limit. Try increasing
max_epochsinoptim_control(), or check if the learning rate is too high. - Alpha distribution WARN: Many tracts have very high values (), meaning they effectively use only the nearest station. This can happen when travel time data has poor coverage.
- Pop-voter gap WARN/FAIL: The total census population differs substantially from the total registered voters. This is expected to some degree (not everyone registers, some register outside their residence), but gaps > 20% suggest data quality issues.
- Unreachable pairs FAIL: Many tract-station pairs have travel times at the ceiling. This means the routing engine couldn’t find paths — check OSM road network coverage.
- Alpha spatial CV WARN: Alpha values have very low variation across tracts (CV < 0.1). The optimizer may not be adapting to local conditions, possibly because of insufficient data or poor convergence.
2. Optimization Convergence: plot_convergence()
Intuition
The optimizer minimizes Poisson deviance by adjusting the decay
parameters
.
plot_convergence() shows the optimization trajectory: is
the loss decreasing smoothly? Is the gradient approaching zero? What
learning rate schedule was used?
Formal definition
- Loss: Poisson deviance , where = observed census population and = fitted population from the model.
- Gradient norm: , the Euclidean norm of the gradient of the loss with respect to all alpha parameters.
- Learning rate: Step size in gradient descent. May decay over epochs via cosine annealing or other schedules.
Applied examples
All three panels (loss, gradient, learning rate):
plot_convergence(result_vga)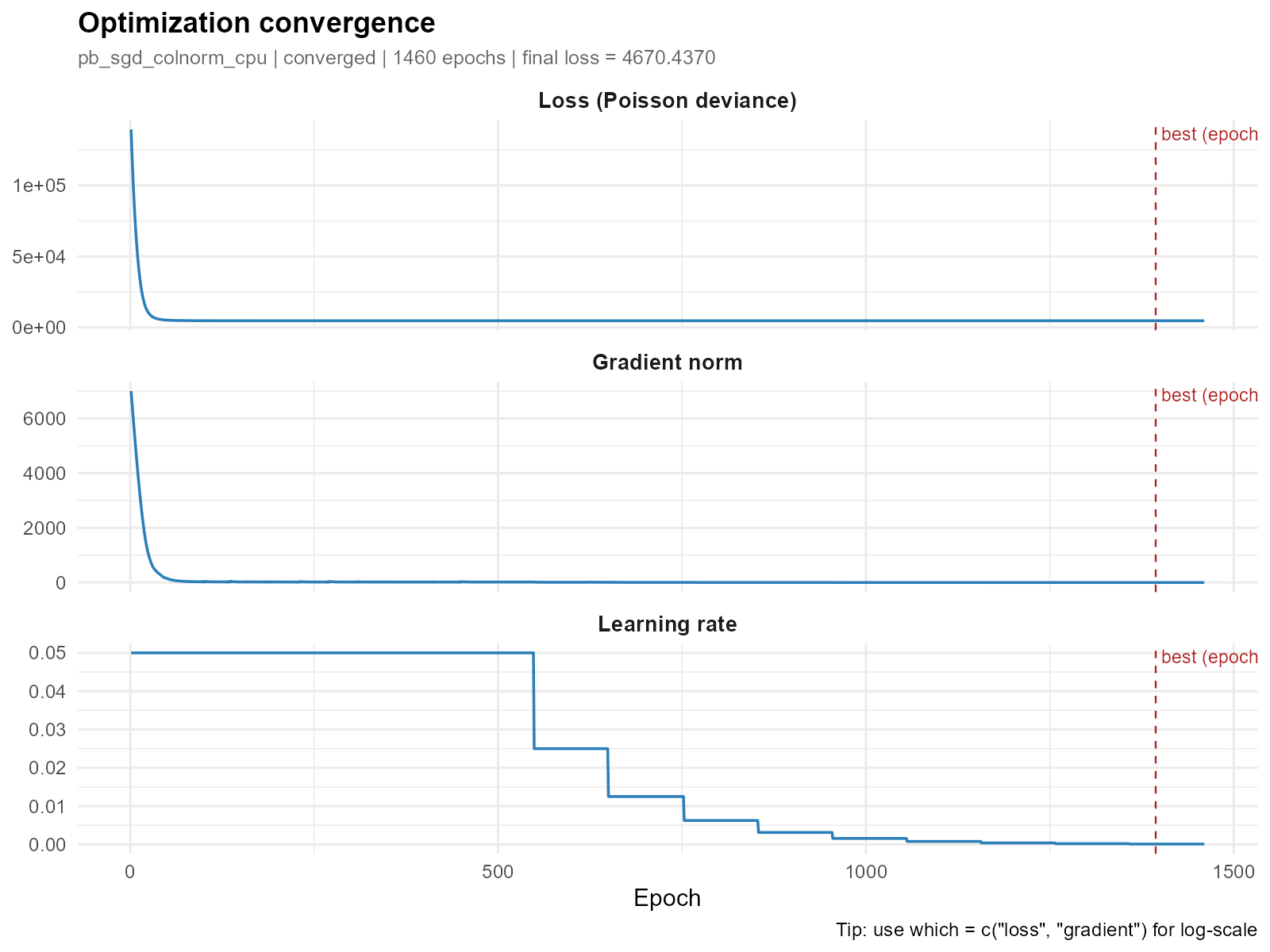
Comparing with a GPU-optimized large city:
plot_convergence(result_bh)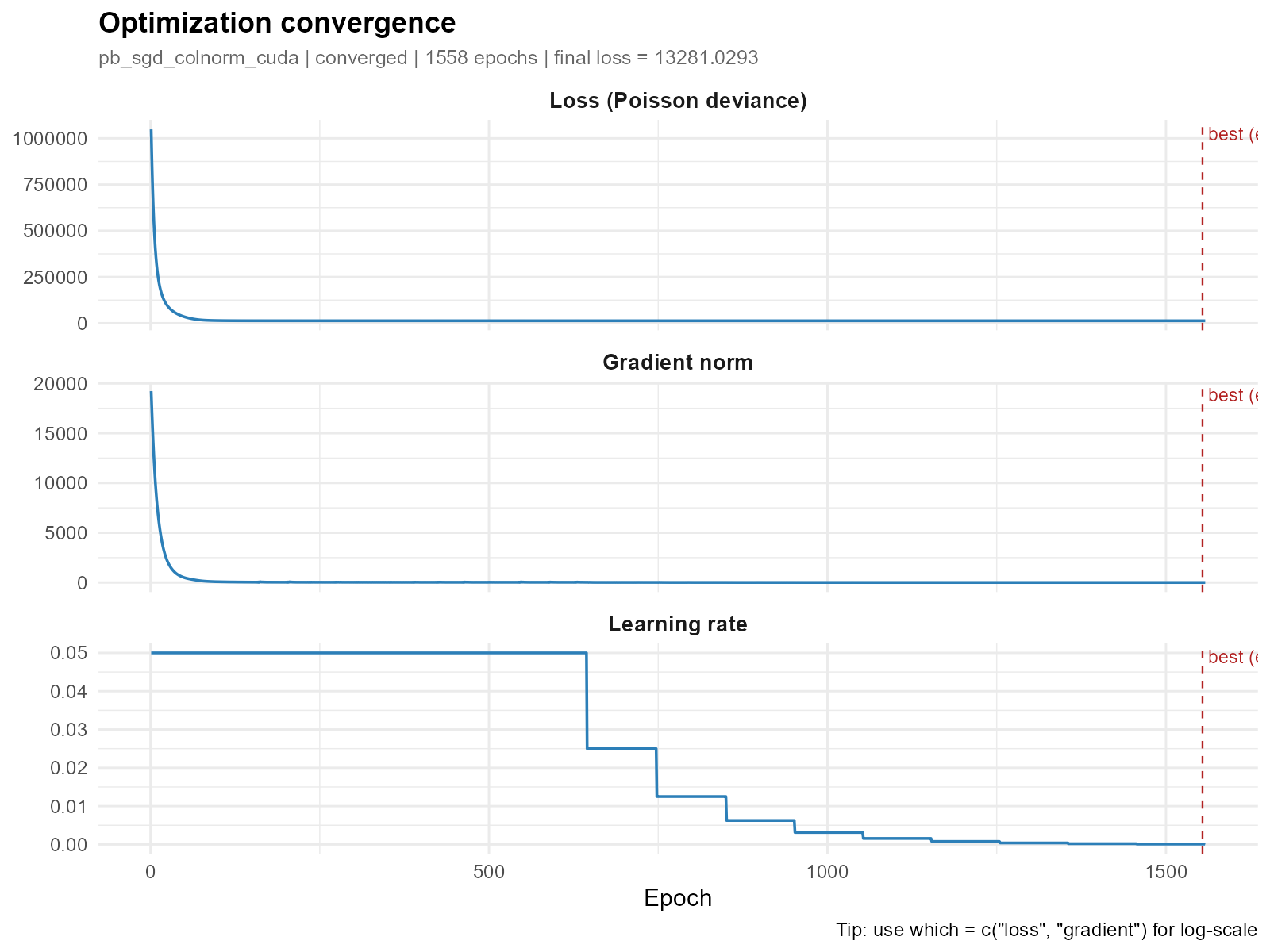
Show only the loss panel:
plot_convergence(result_vga, which = "loss")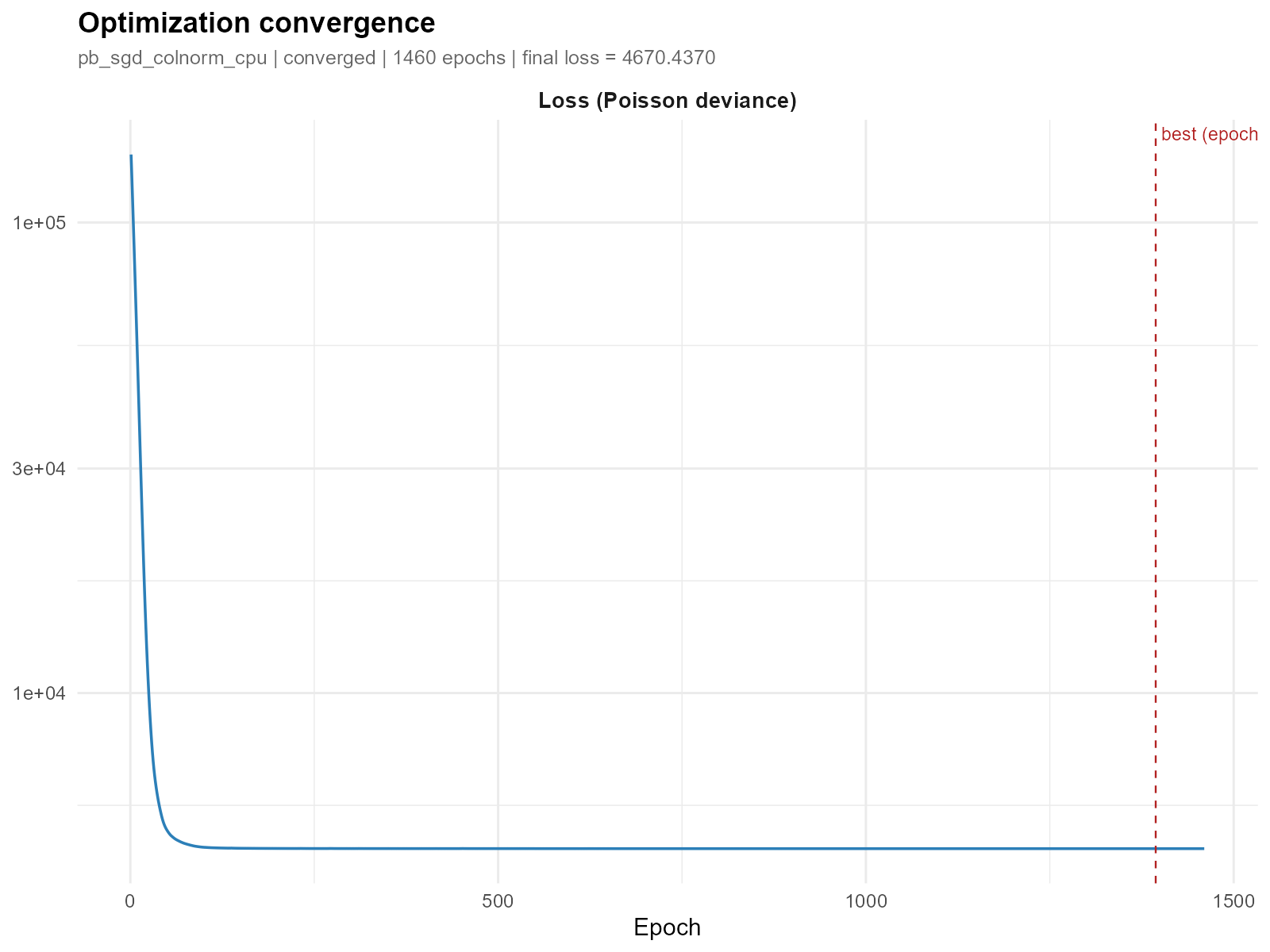
Log scale for loss and gradient (useful when the values span orders of magnitude):
plot_convergence(result_vga, which = c("loss", "gradient"), log_y = TRUE)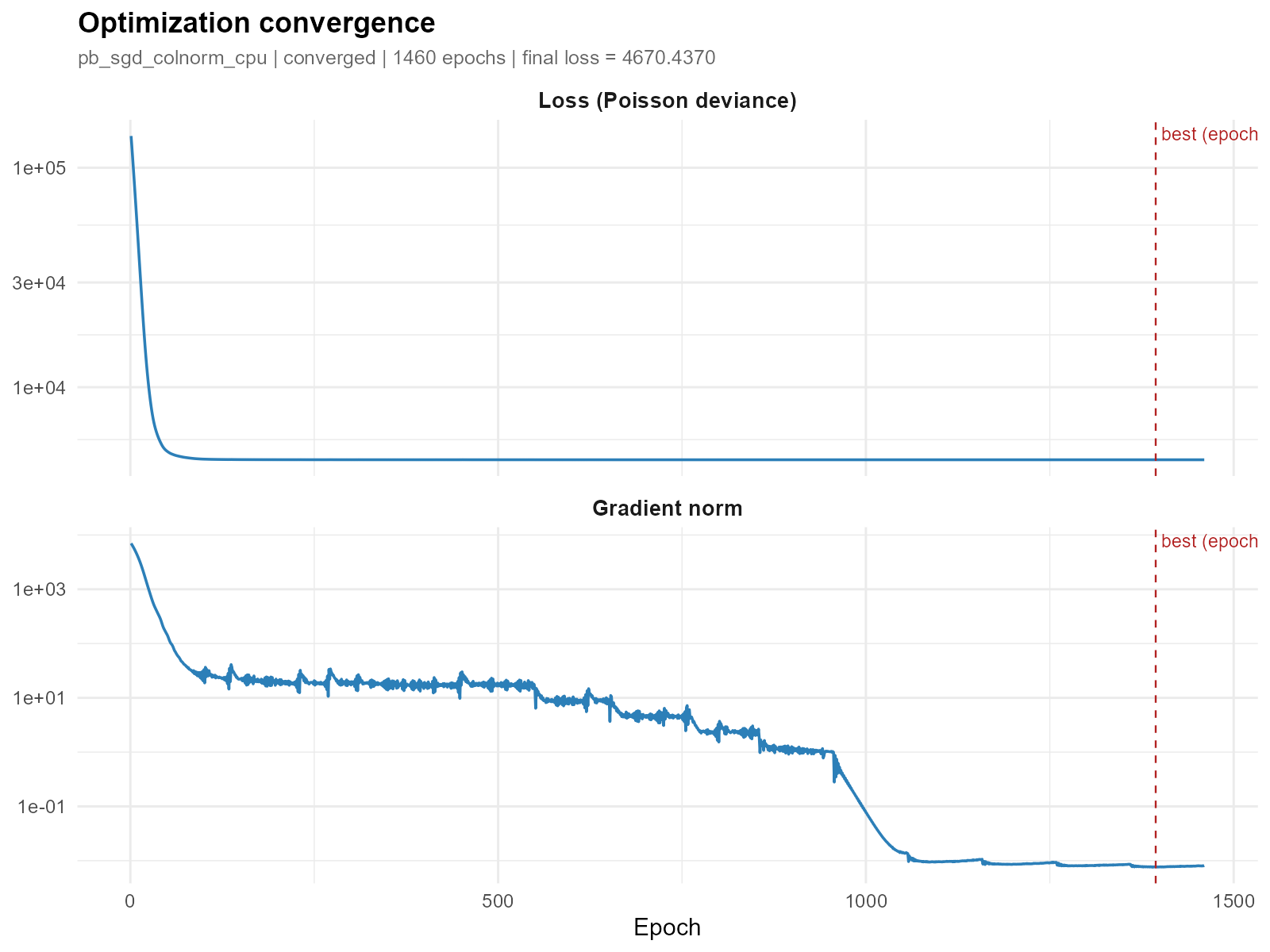
Interpretation
- Good convergence: Smooth, monotonically decreasing loss that plateaus. Gradient norm approaches zero. The vertical dashed red line marks the best epoch.
-
Poor convergence: Oscillating loss, gradient that
doesn’t shrink, or loss that hasn’t plateaued by the last epoch.
Consider increasing
max_epochsor adjusting the learning rate. - Learning rate: The default cosine annealing with warm restarts produces a sawtooth pattern — LR decays to a floor then jumps back up at each restart cycle.
3. Alpha Decay Parameters: plot_alpha()
Intuition
The parameter controls how sharply the weight of station decays with travel time for tract . Higher means voters are more localized (sharp decay — only nearby stations matter). Lower means voters come from a broader area (gradual decay — distant stations still contribute).
is a matrix with one row per tract and one column per demographic bracket (14 brackets = 7 age groups 2 genders). Different brackets may have different decay patterns (e.g., young voters may travel farther to vote).
Formal definition
The weight before column normalization is:
For the default power-law kernel:
For the exponential kernel
(optim_control(kernel = "exponential")):
The final weight is
(column-normalized so each station’s voters are fully distributed). See
vignette("methodology") for details on kernel choice.
Applied examples: Map mode
The map mode collapses the 14-bracket alpha matrix to a single scalar
per tract using summary_fn. This is the key visualization
choice — different summaries answer different questions.
Median (default) — the “typical” alpha across all brackets:
plot_alpha(result_vga, type = "map", summary_fn = "median")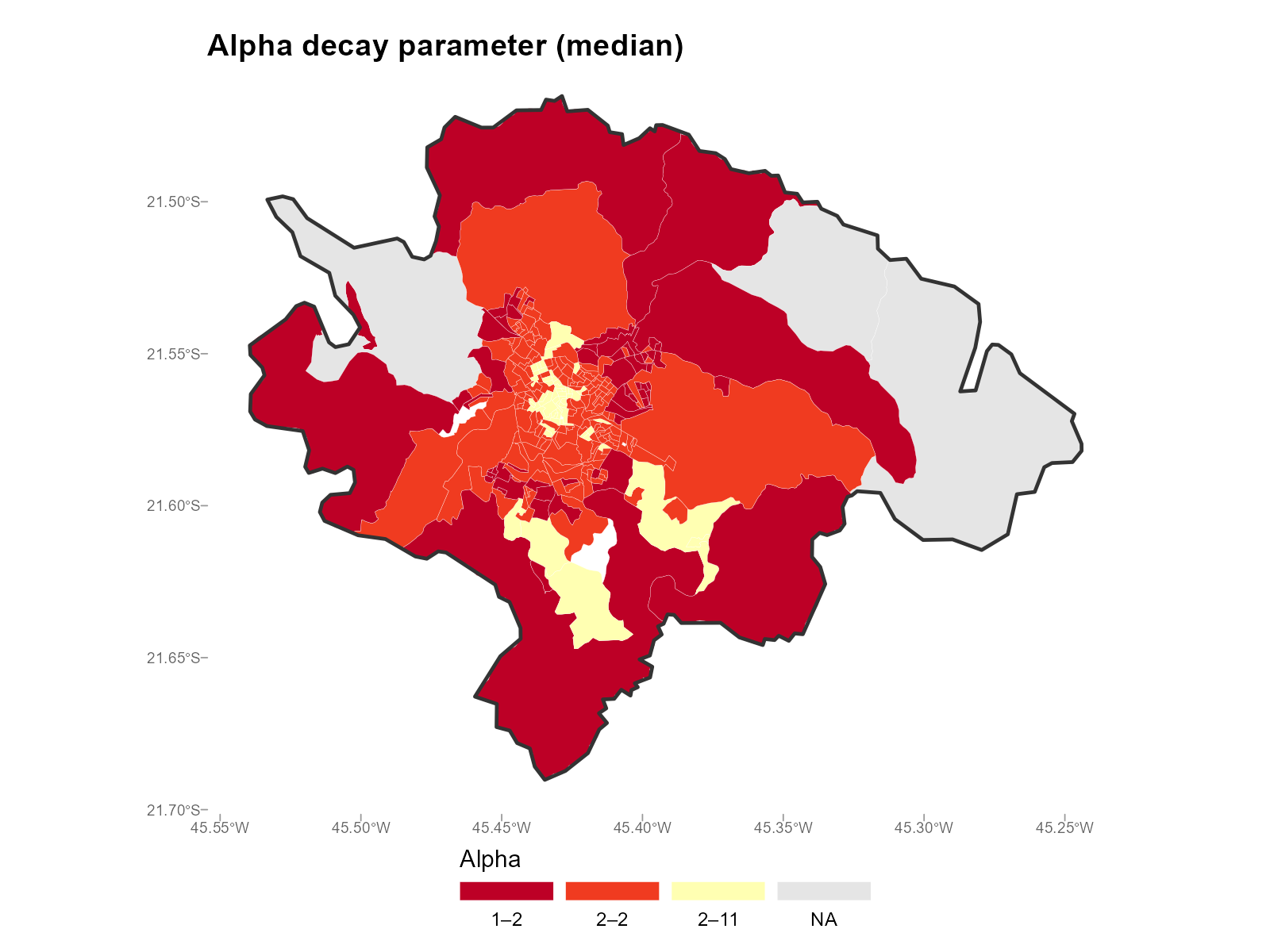
Mean — sensitive to extreme values in any bracket:
plot_alpha(result_vga, type = "map", summary_fn = "mean")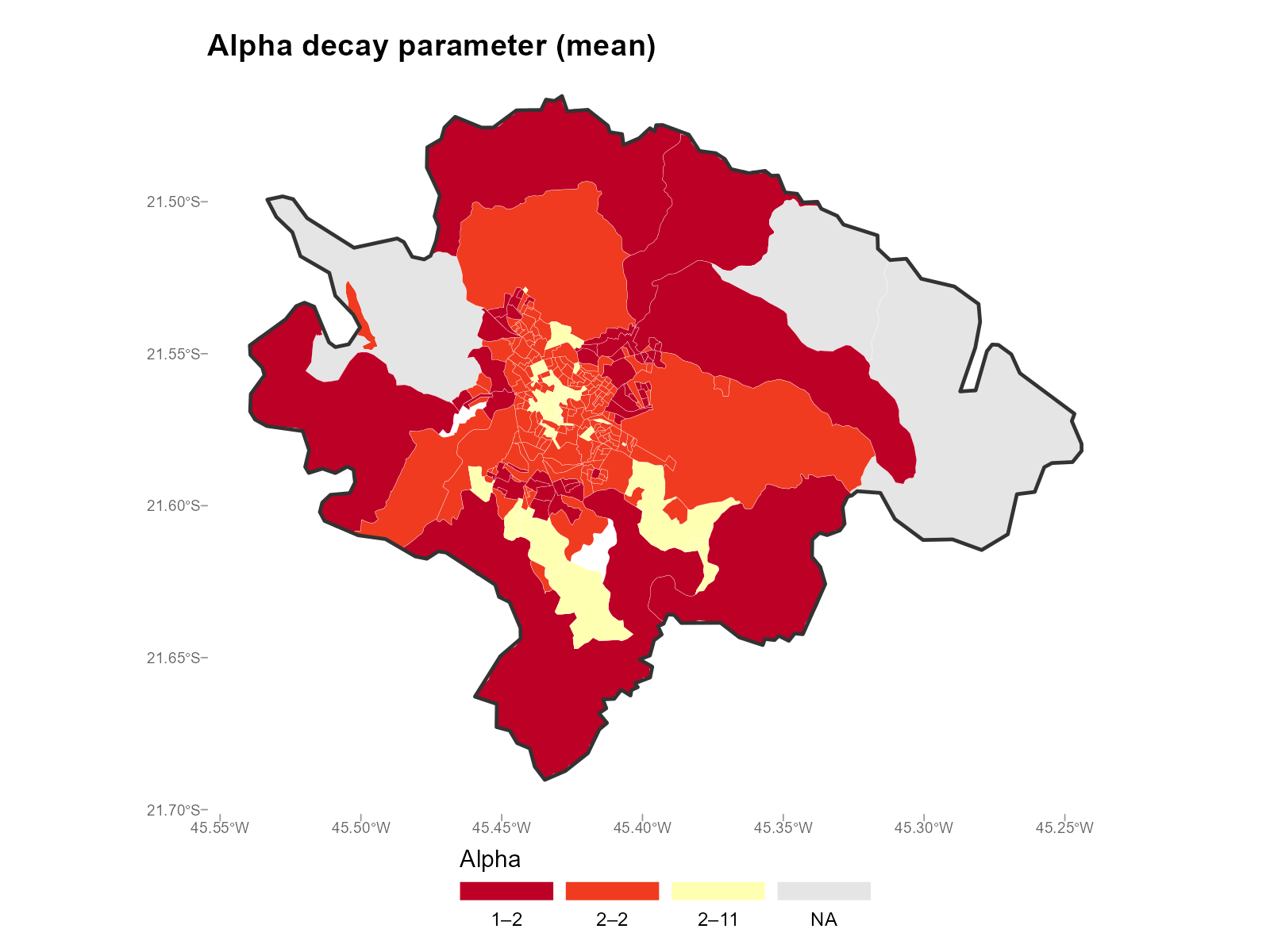
Population-weighted — weights each bracket by the tract’s population in that bracket:
plot_alpha(result_vga, type = "map", summary_fn = "pop_weighted")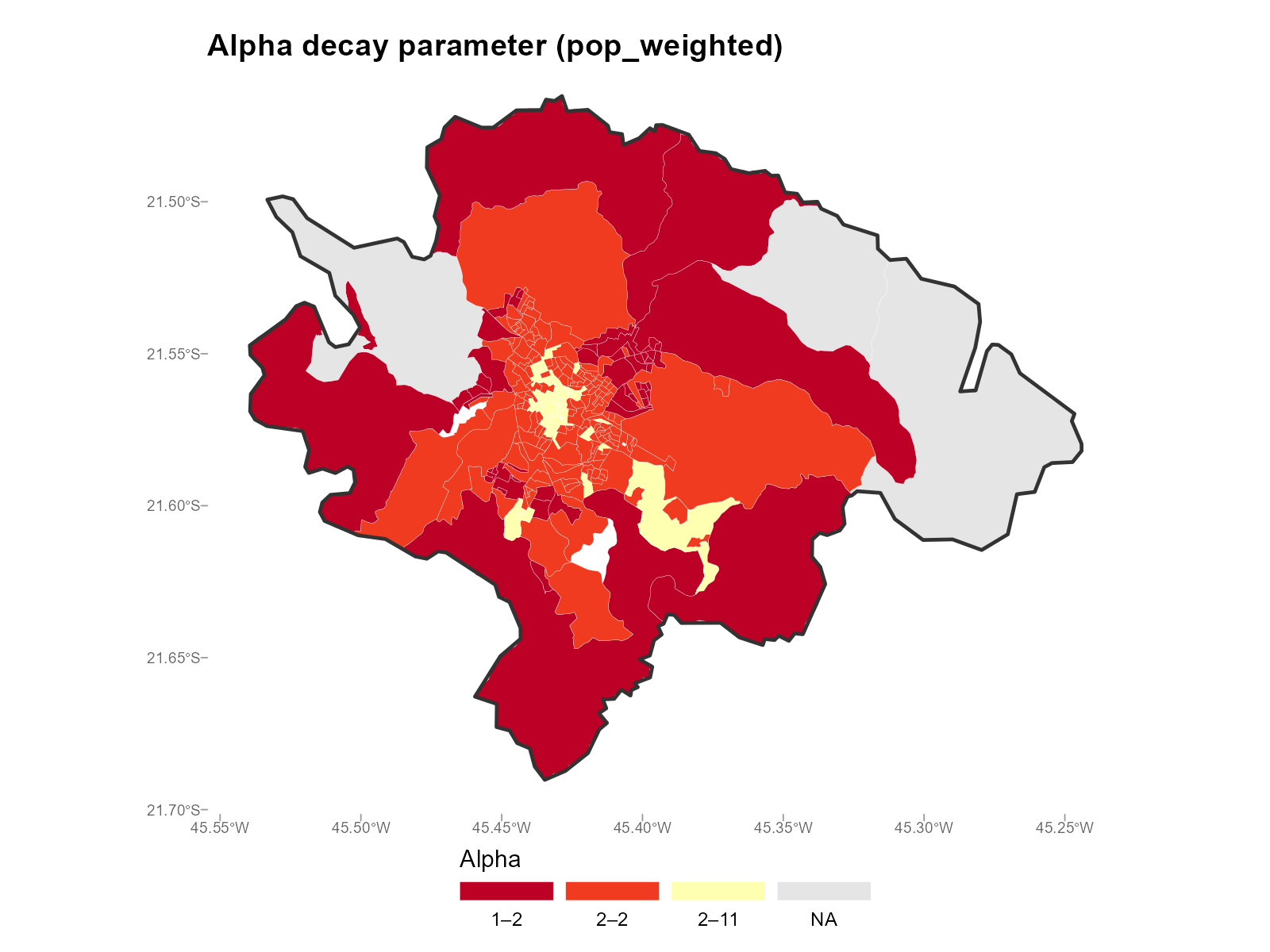
Range () — measures how much varies across brackets within each tract:
plot_alpha(result_vga, type = "map", summary_fn = "range")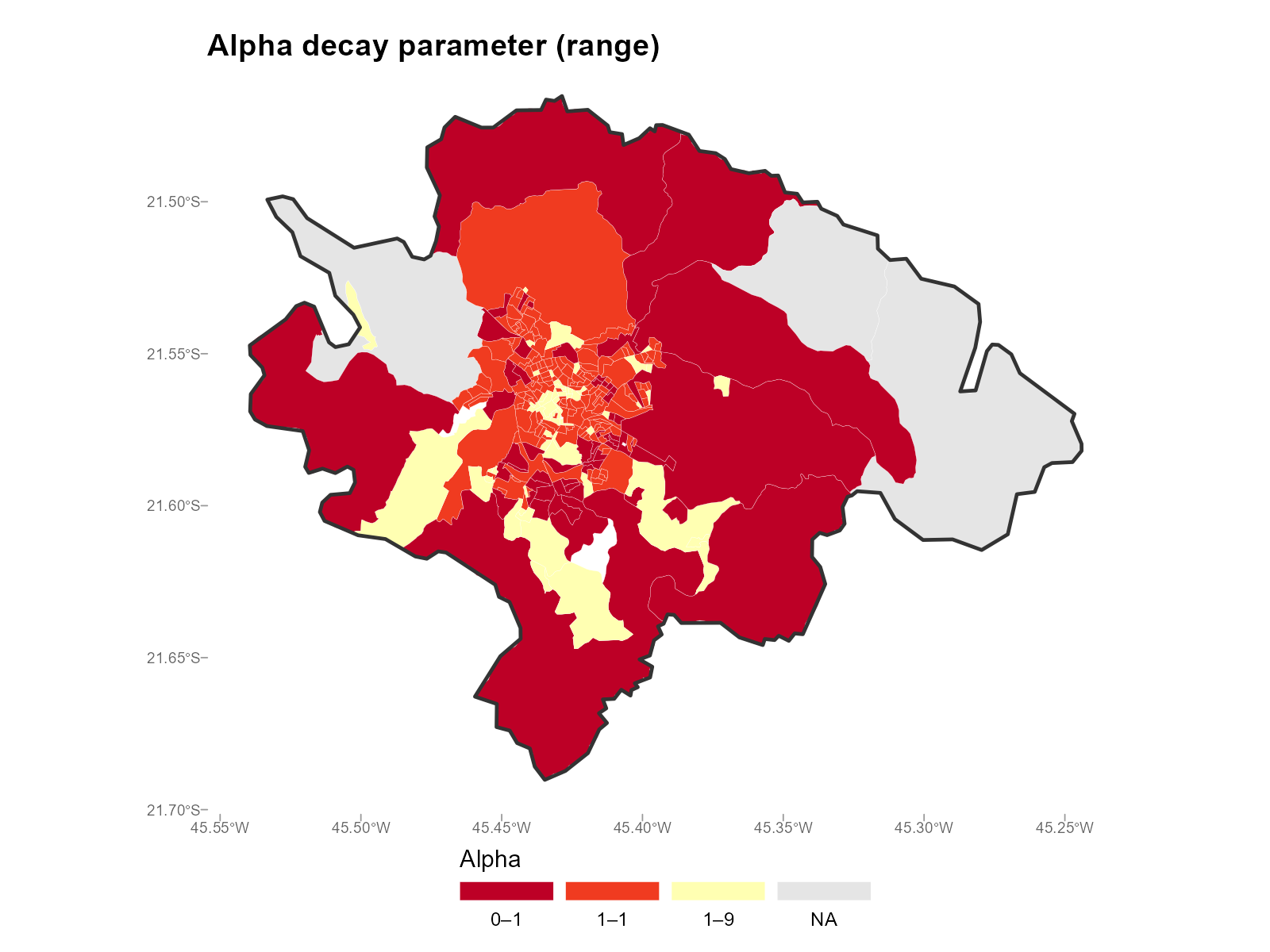
Single bracket (e.g., bracket 1 = Male 18–20):
plot_alpha(result_vga, type = "map", summary_fn = 1)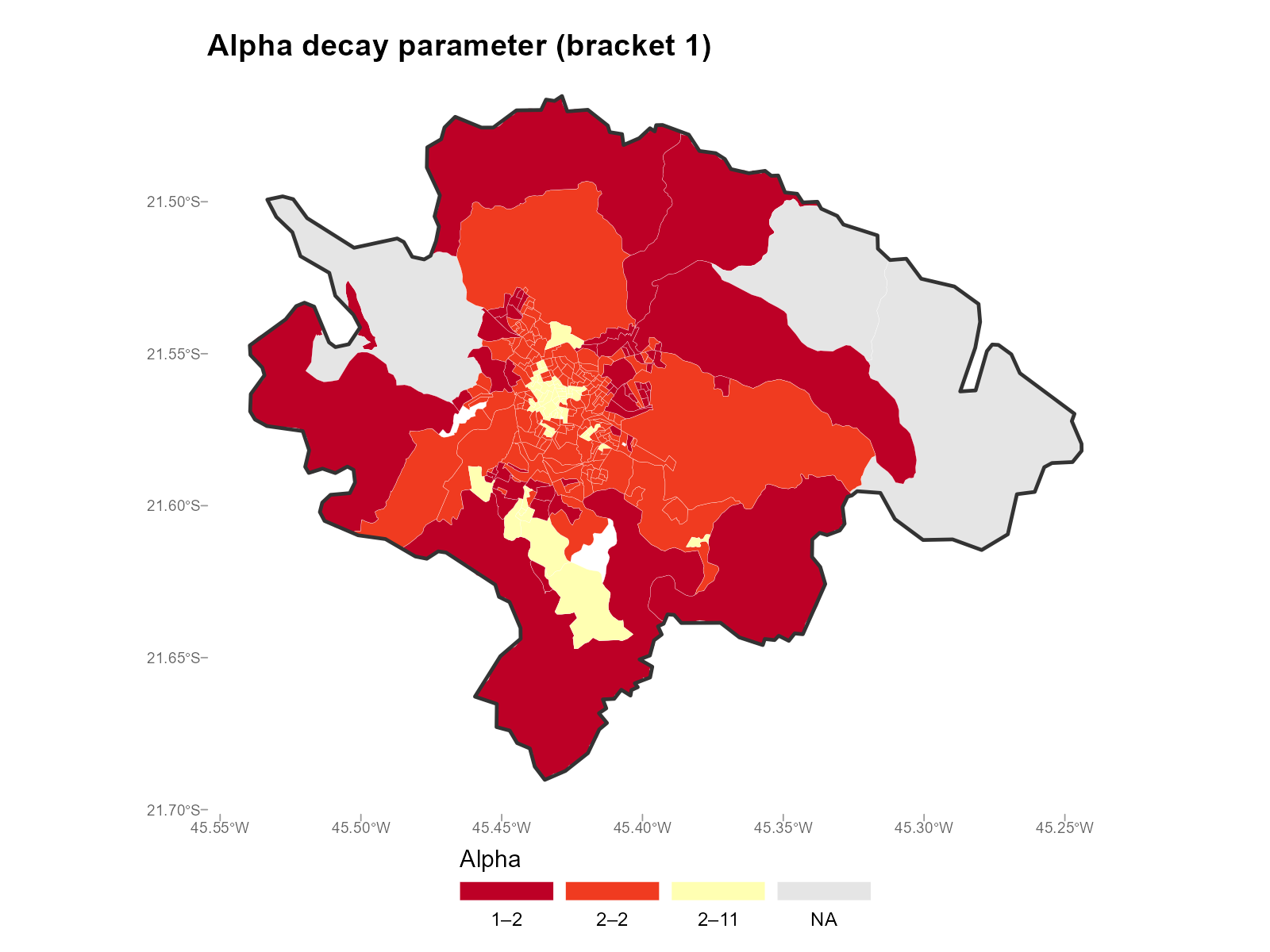
Large city comparison (Belo Horizonte):
plot_alpha(result_bh, type = "map", summary_fn = "median")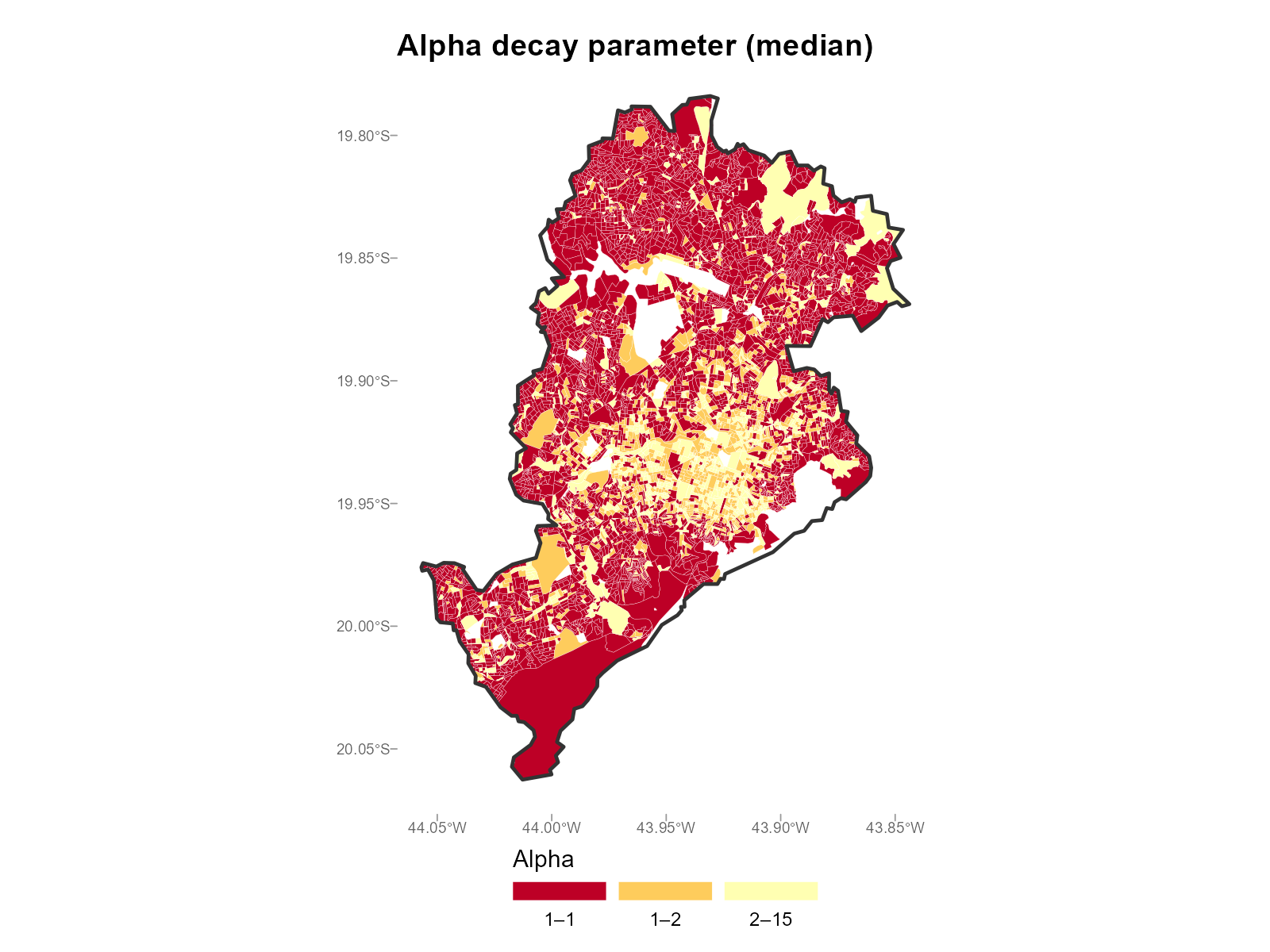
Applied examples: Histogram mode
Density plots faceted by bracket. With 14 brackets arranged in a 2$$7 grid, you can compare age groups and genders at a glance.
plot_alpha(result_nit, type = "histogram")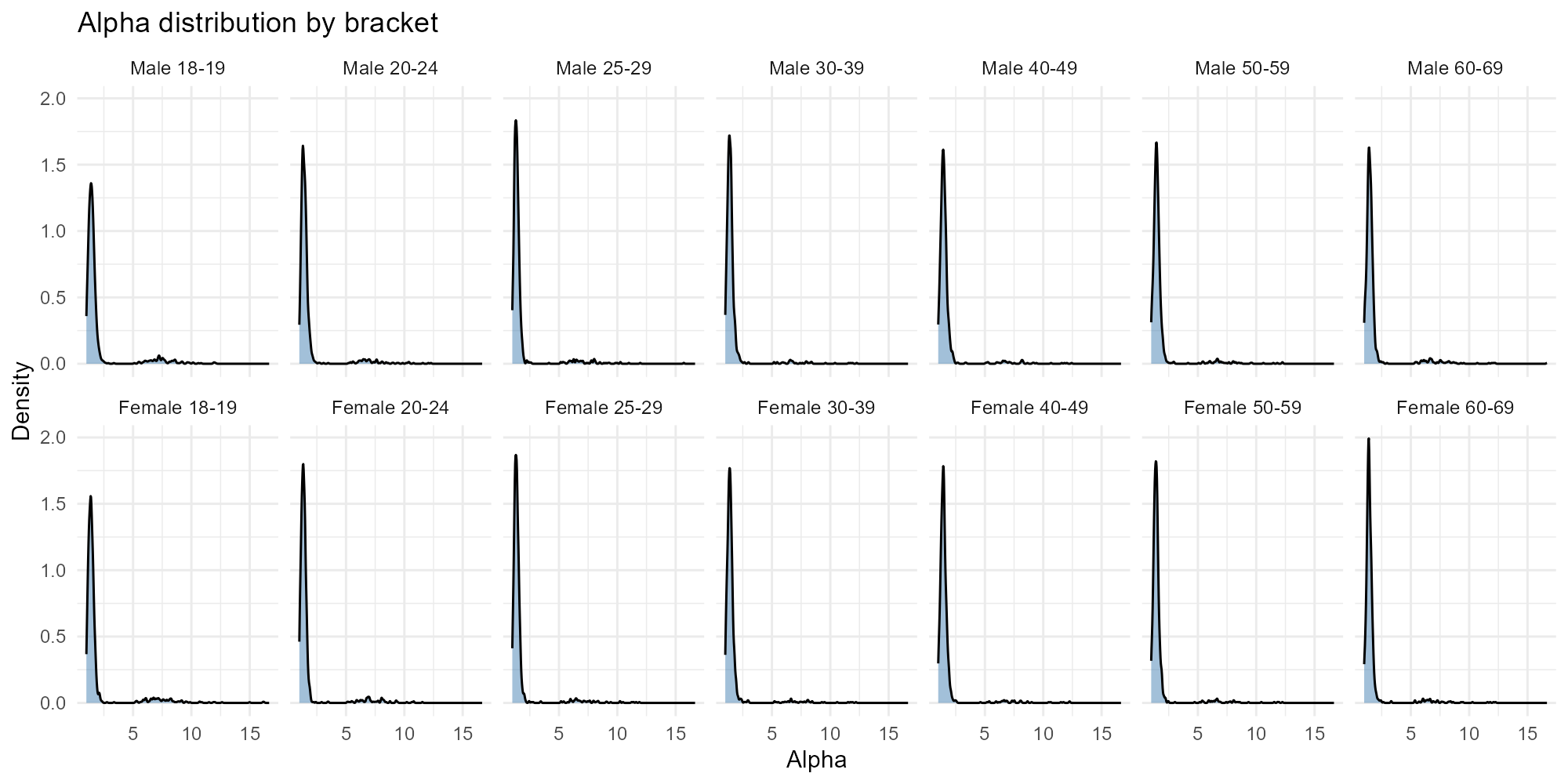
Select specific brackets:
plot_alpha(result_nit, type = "histogram", brackets = c(1, 7, 8, 14))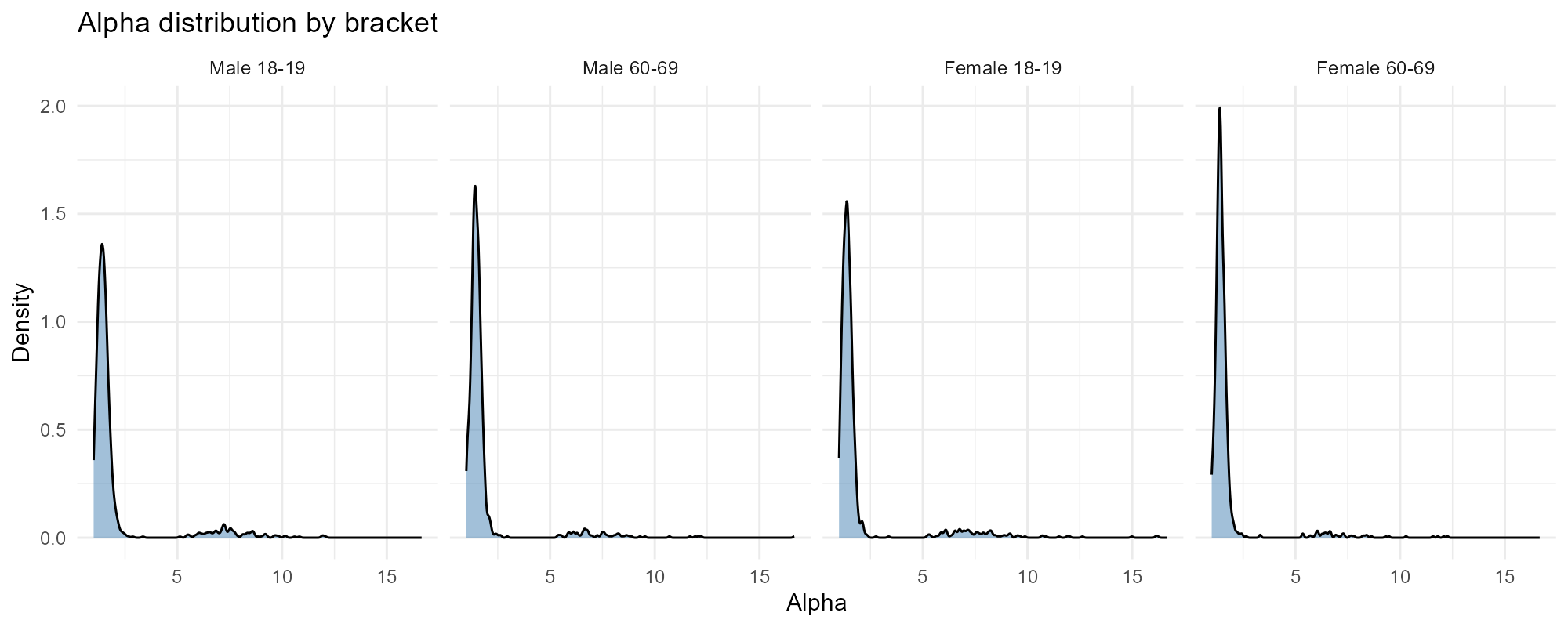
Small city (narrower distributions due to fewer tracts):
plot_alpha(result_igr, type = "histogram")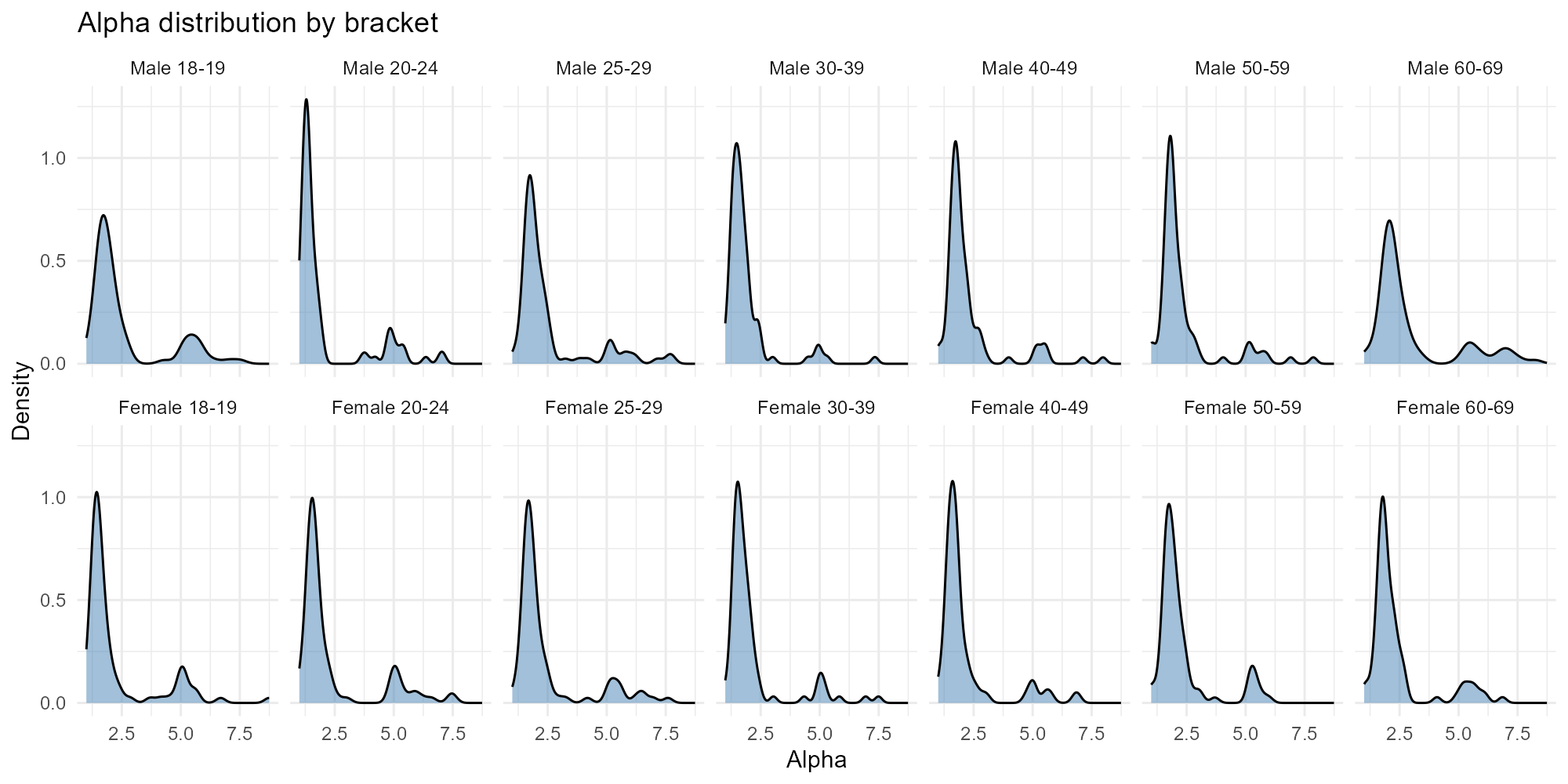
Applied examples: Bracket boxplot
The most compact view — one boxplot per bracket on a single panel:
plot_alpha(result_bh, type = "bracket")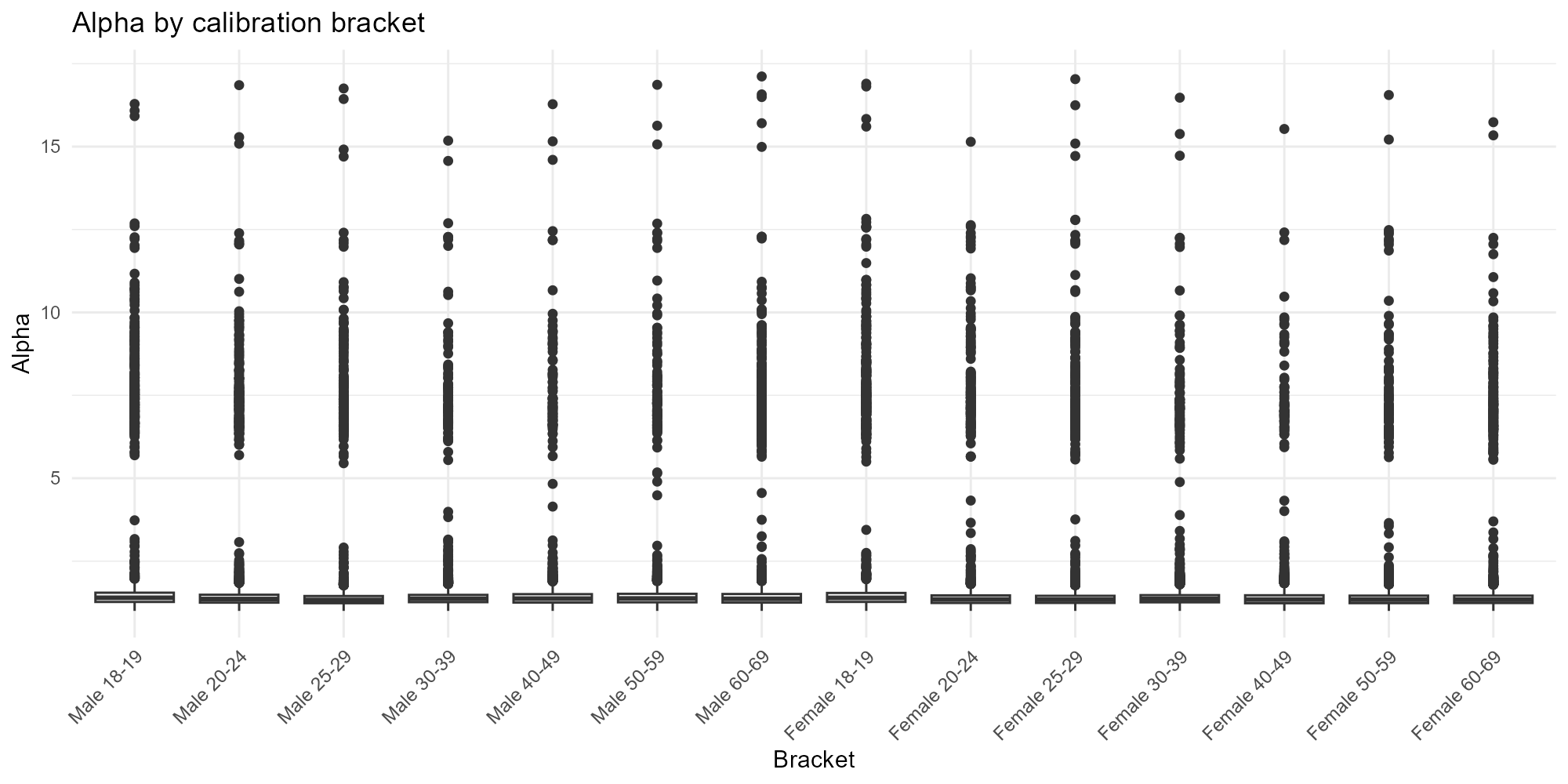
Male brackets only:
plot_alpha(result_nit, type = "bracket", brackets = 1:7)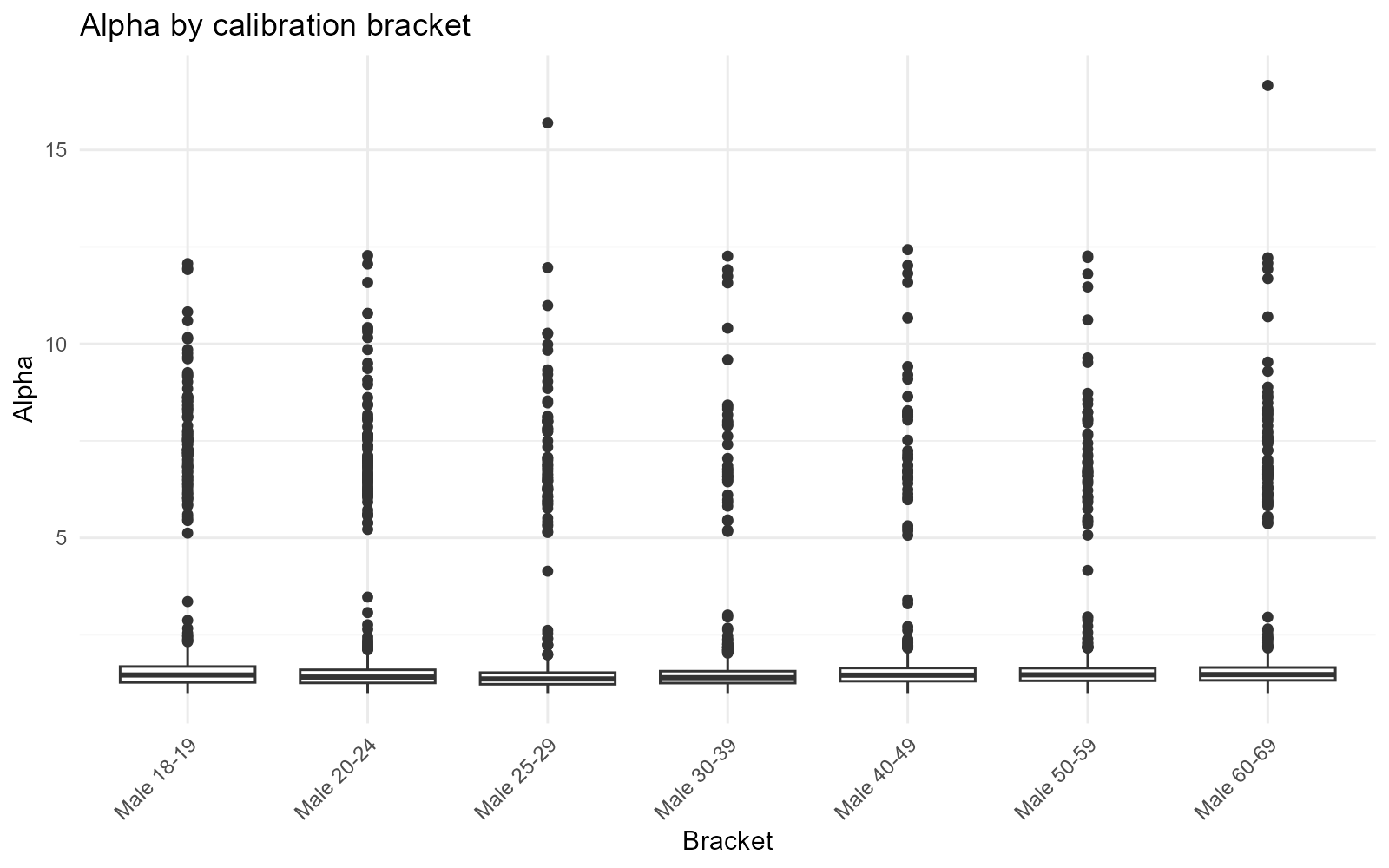
Interpretation
- Urban core vs periphery: Central tracts with many nearby stations tend to have higher (sharper decay). Peripheral tracts with few nearby stations have lower (they must draw from distant stations).
- Age patterns: Younger brackets may show different distributions than older ones, reflecting different mobility patterns.
- Range map: High range = the optimizer found very different decay rates for different brackets in that tract. This often occurs at the urban-rural boundary.
- : Effectively nearest-station assignment. The decay is so steep that only the closest station has non-negligible weight.
4. Calibration Residuals: plot_residuals() +
residual_summary()
Intuition
The model calibrates against known census demographics (population by agegender bracket). Residuals measure how well the model reproduces these known quantities. Small residuals mean the model’s spatial allocation of voters closely matches census demographics — giving confidence that the interpolated (unknown) electoral variables are also spatially accurate.
Formal definition
Three types of residuals are available:
- Raw: (fitted minus observed). Scale depends on population size.
- Pearson: . Standardized under the Poisson assumption — comparable across tracts of different sizes.
- Deviance: . Individual Poisson deviance contributions.
Applied examples: Map mode
For maps, summary_fn collapses across the 14 brackets to
a single value per tract.
RMSE (default) — magnitude of misfit:
plot_residuals(result_vga, type = "map", summary_fn = "rmse")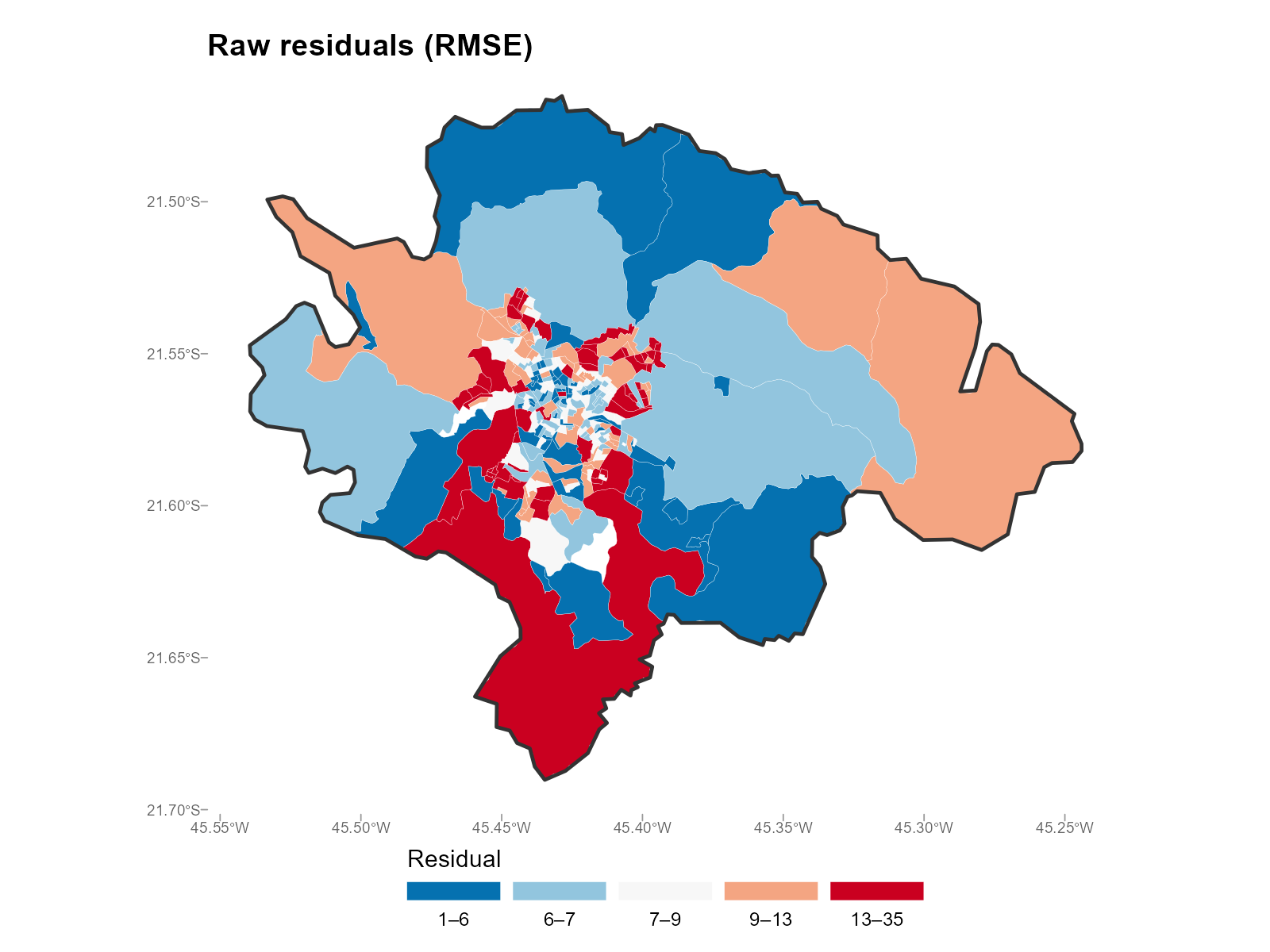
Mean — directional bias (positive = over-prediction, negative = under-prediction):
plot_residuals(result_vga, type = "map", summary_fn = "mean")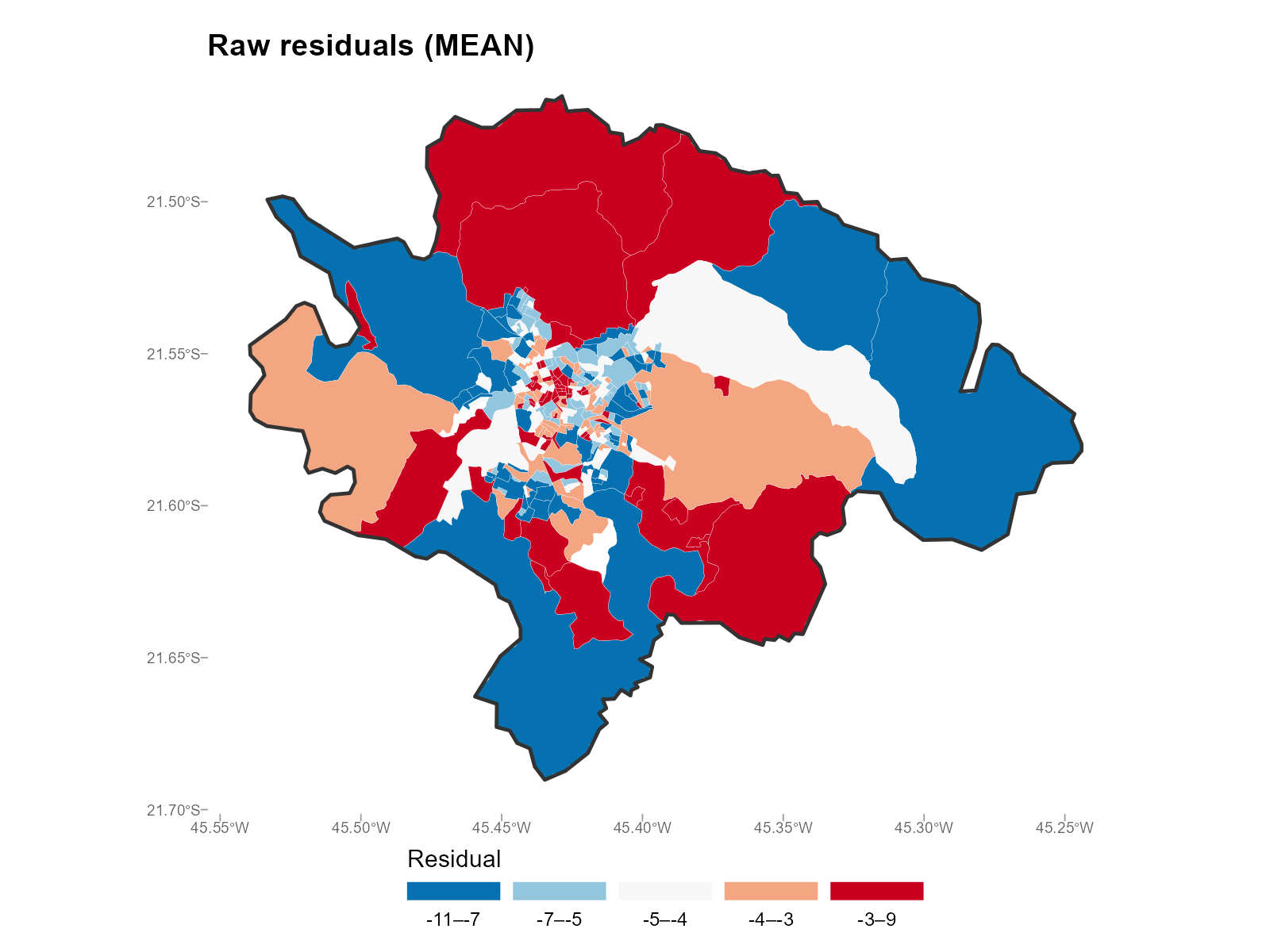
Pearson residuals (standardized):
plot_residuals(result_vga, type = "map", residual_type = "pearson",
summary_fn = "rmse")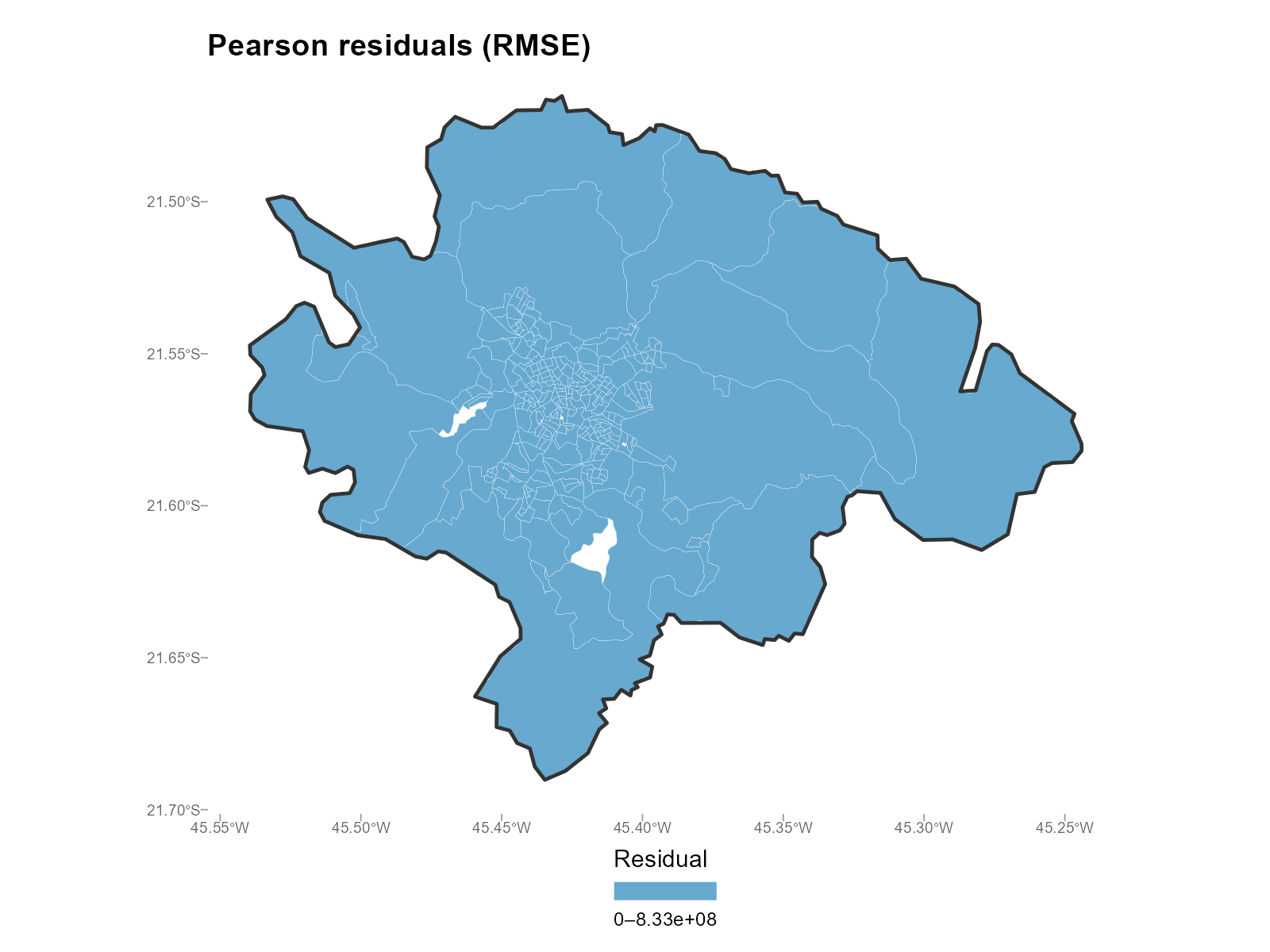
Large city (Belo Horizonte):
plot_residuals(result_bh, type = "map", summary_fn = "rmse")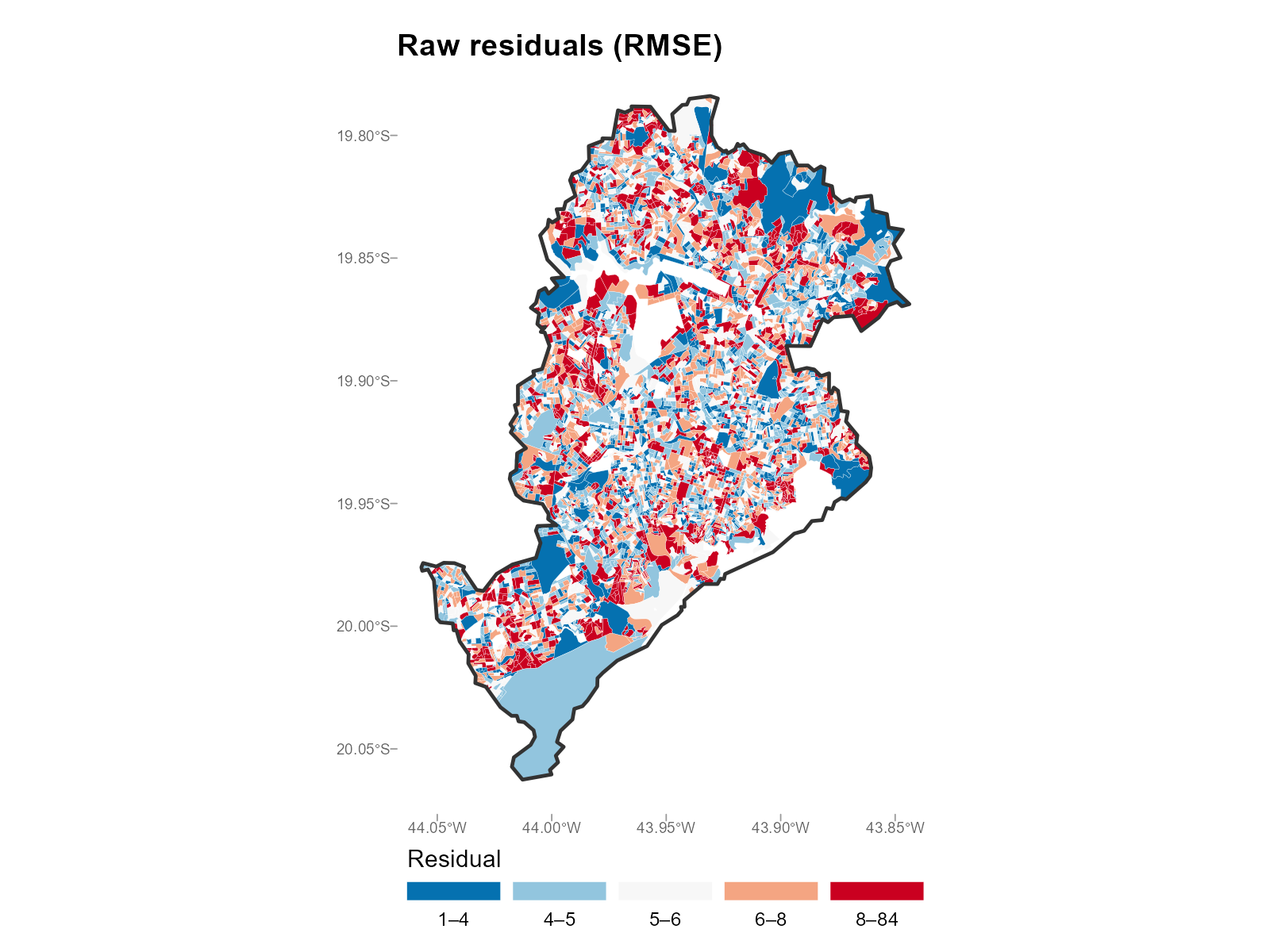
Single bracket:
plot_residuals(result_vga, type = "map", summary_fn = 1)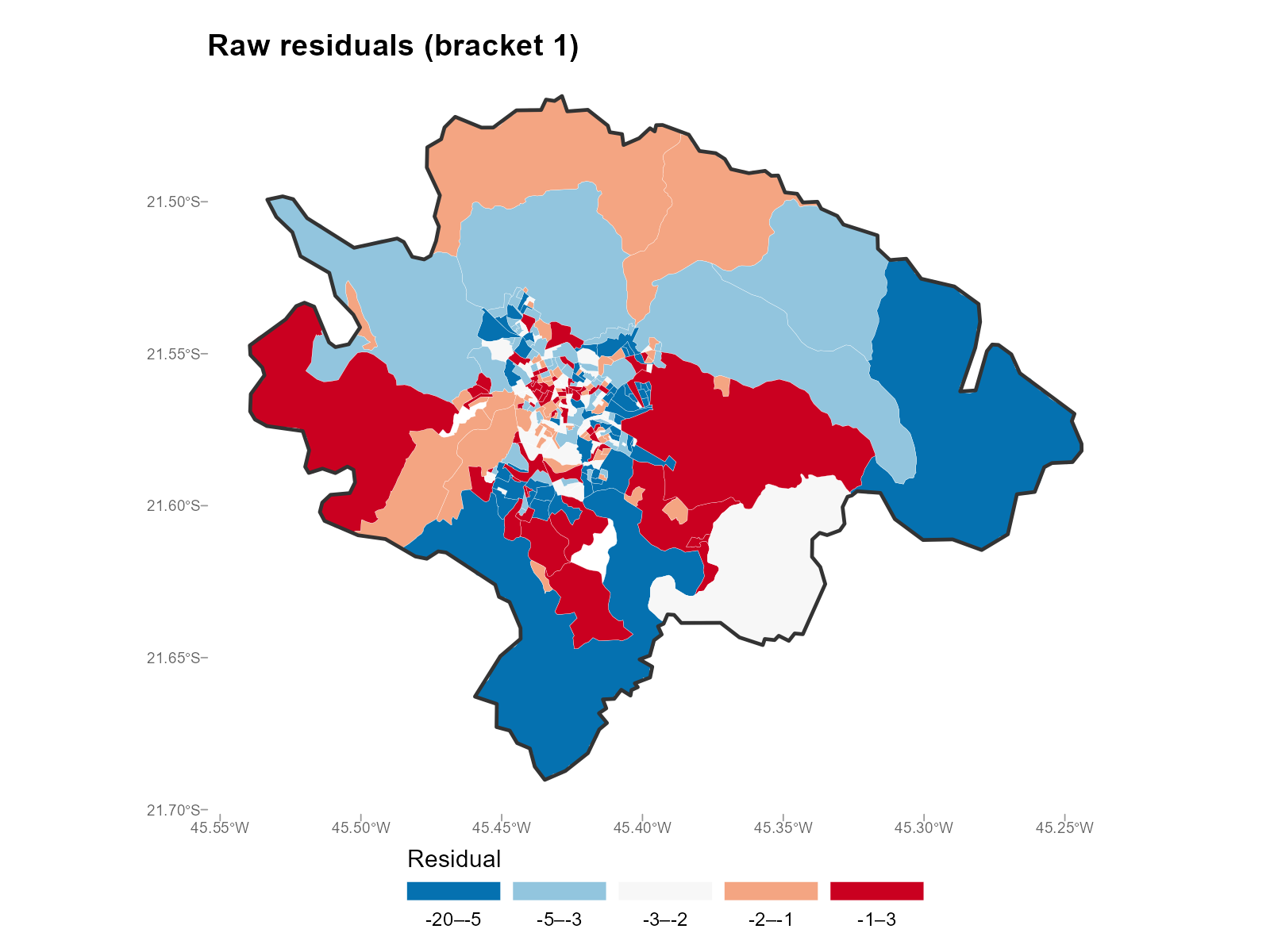
Applied examples: Histogram mode
plot_residuals(result_nit, type = "histogram")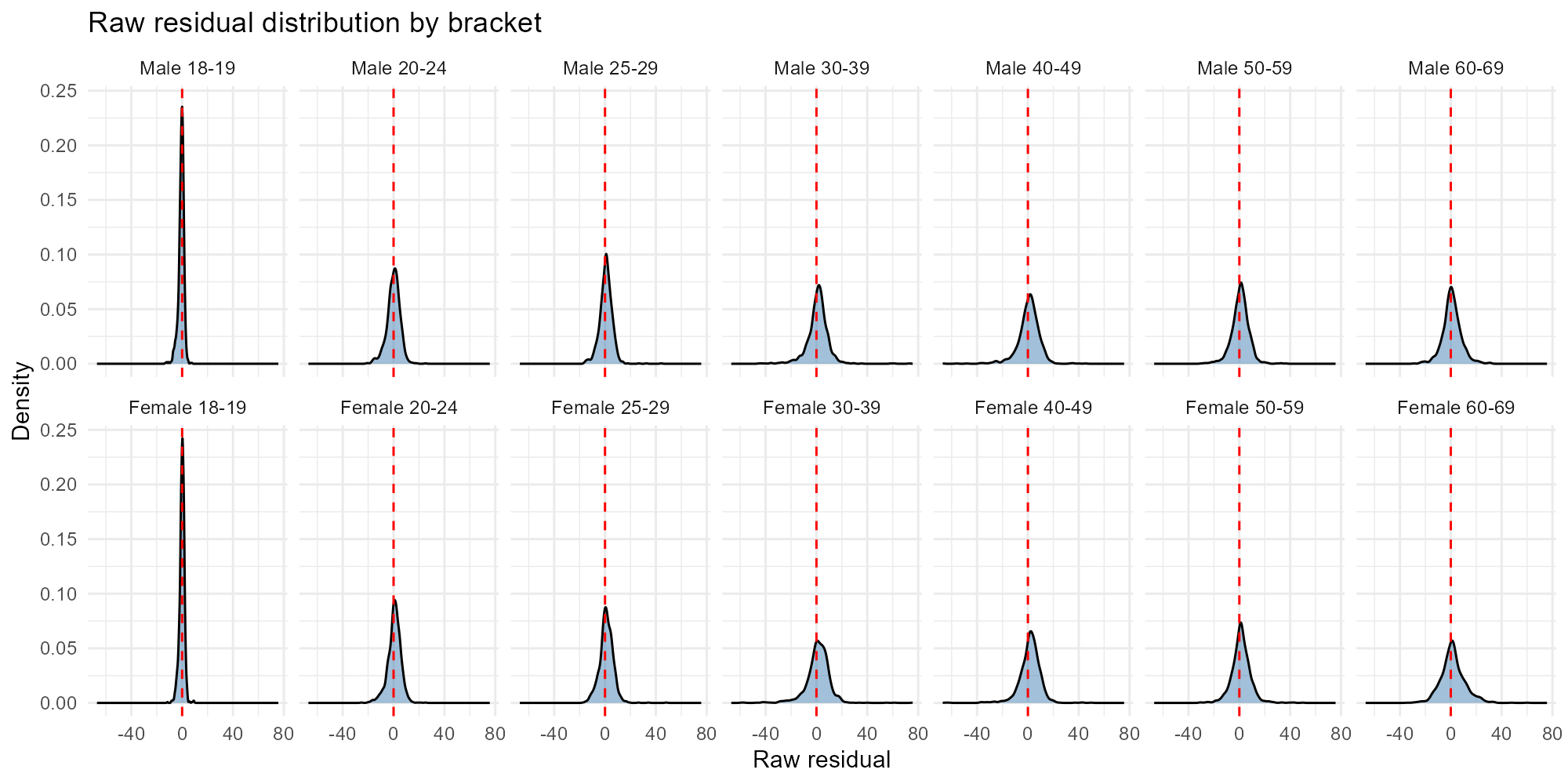
Compare with Pearson residuals:
plot_residuals(result_nit, type = "histogram", residual_type = "pearson")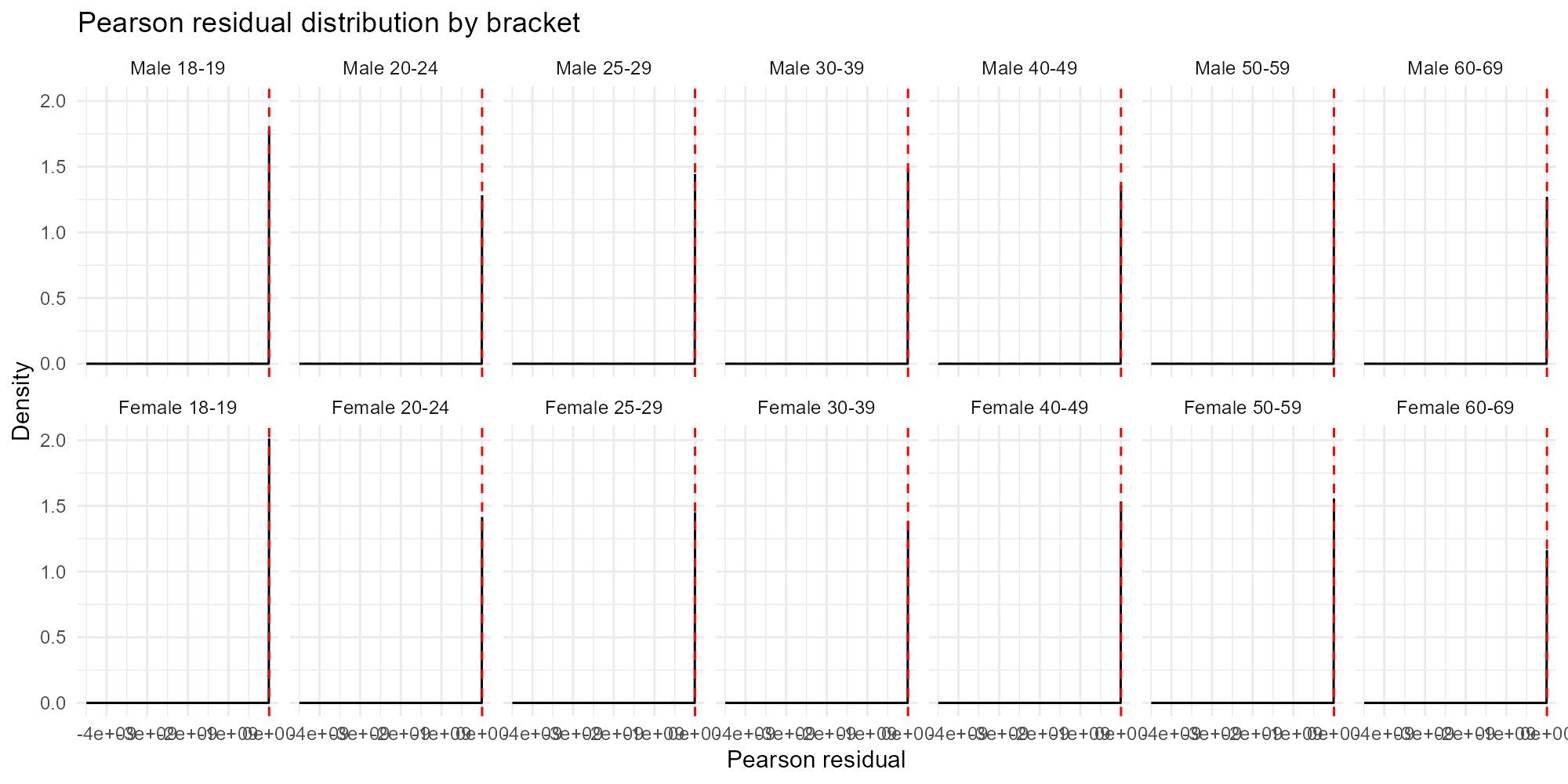
Applied examples: Bracket boxplot
plot_residuals(result_bh, type = "bracket")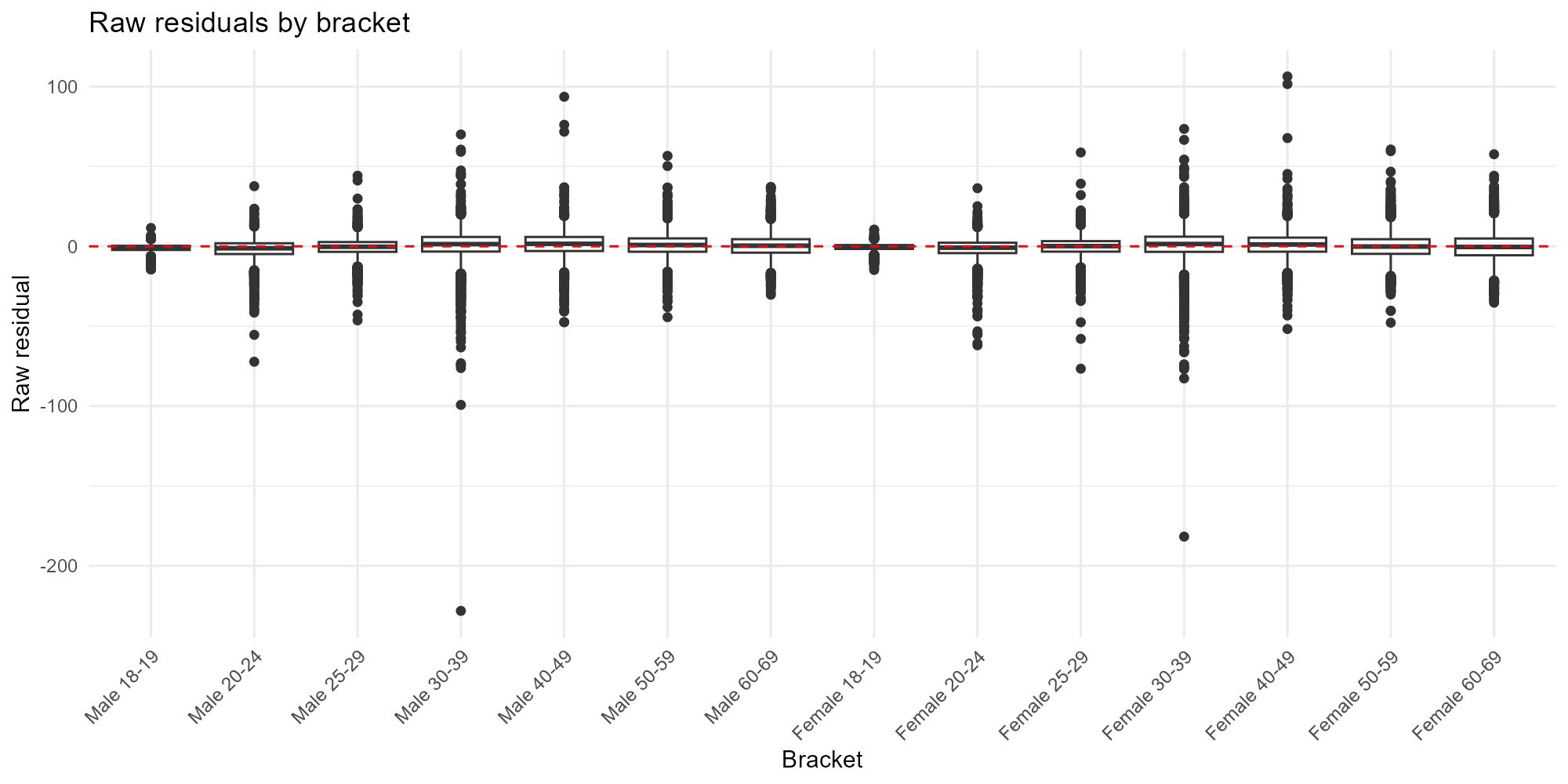
Deviance scale:
plot_residuals(result_bh, type = "bracket", residual_type = "deviance")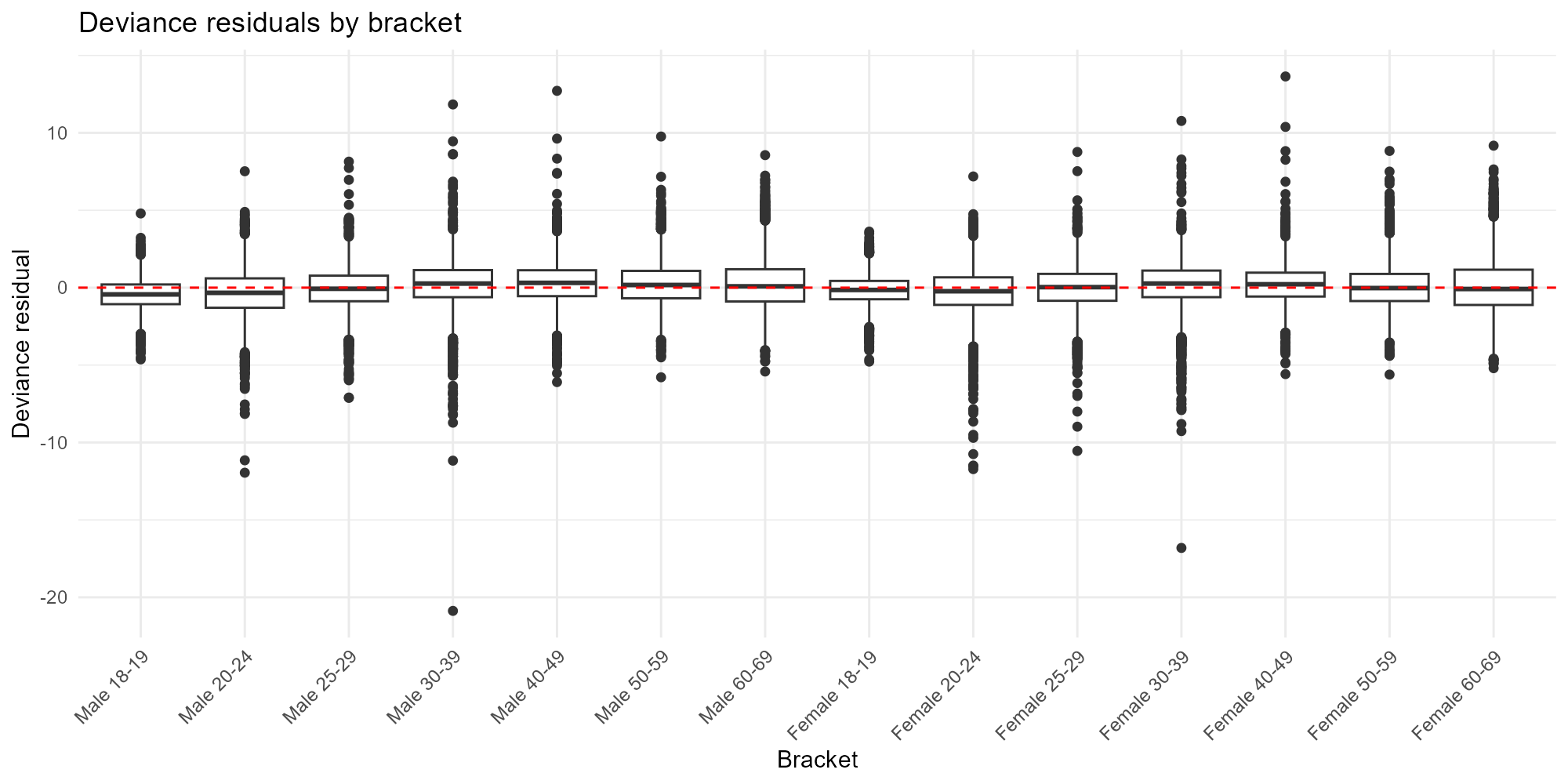
Applied examples: Scatter (fitted vs observed)
plot_residuals(result_igr, type = "scatter")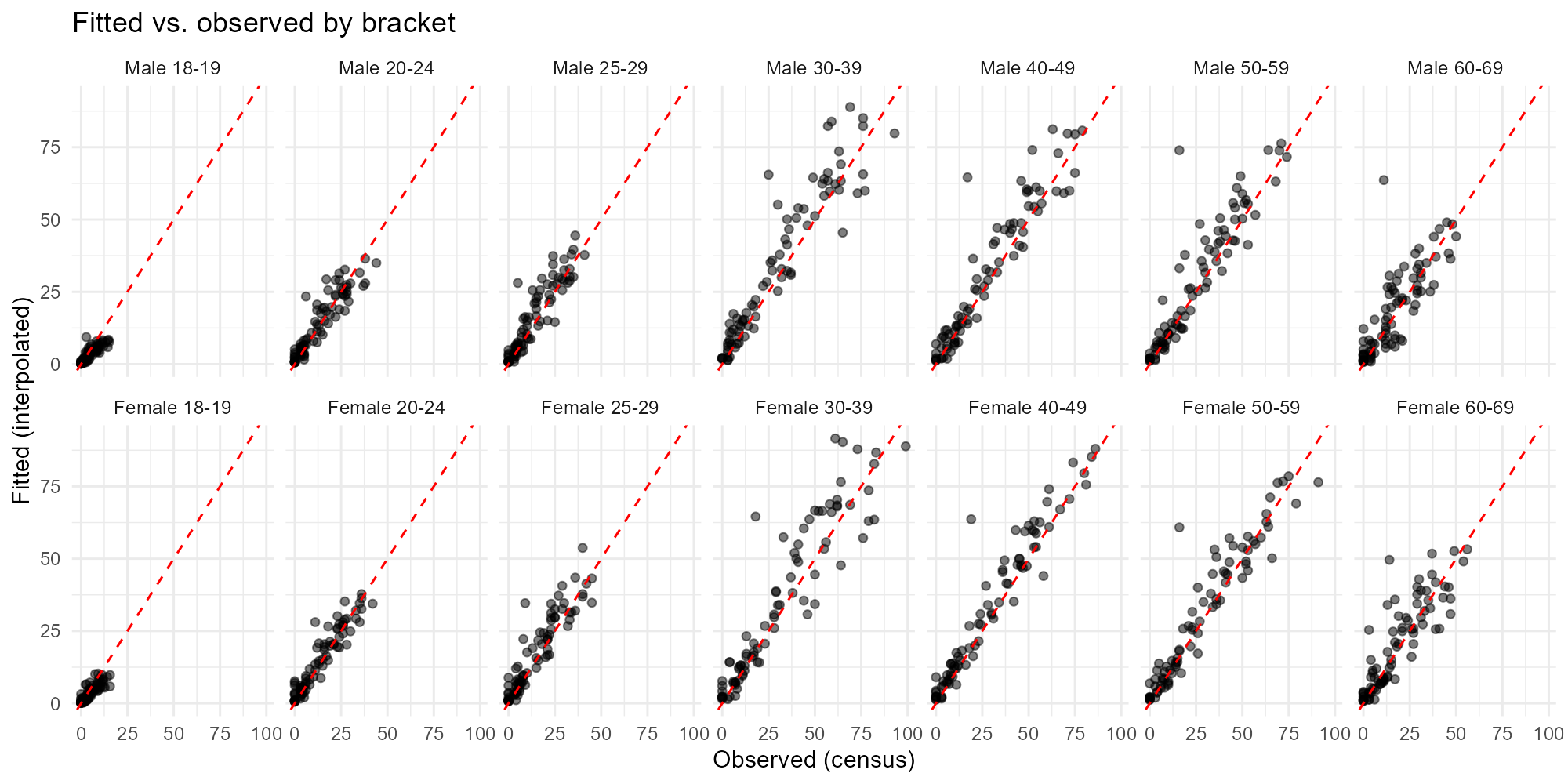
Selected brackets:
plot_residuals(result_vga, type = "scatter", brackets = c(1, 14))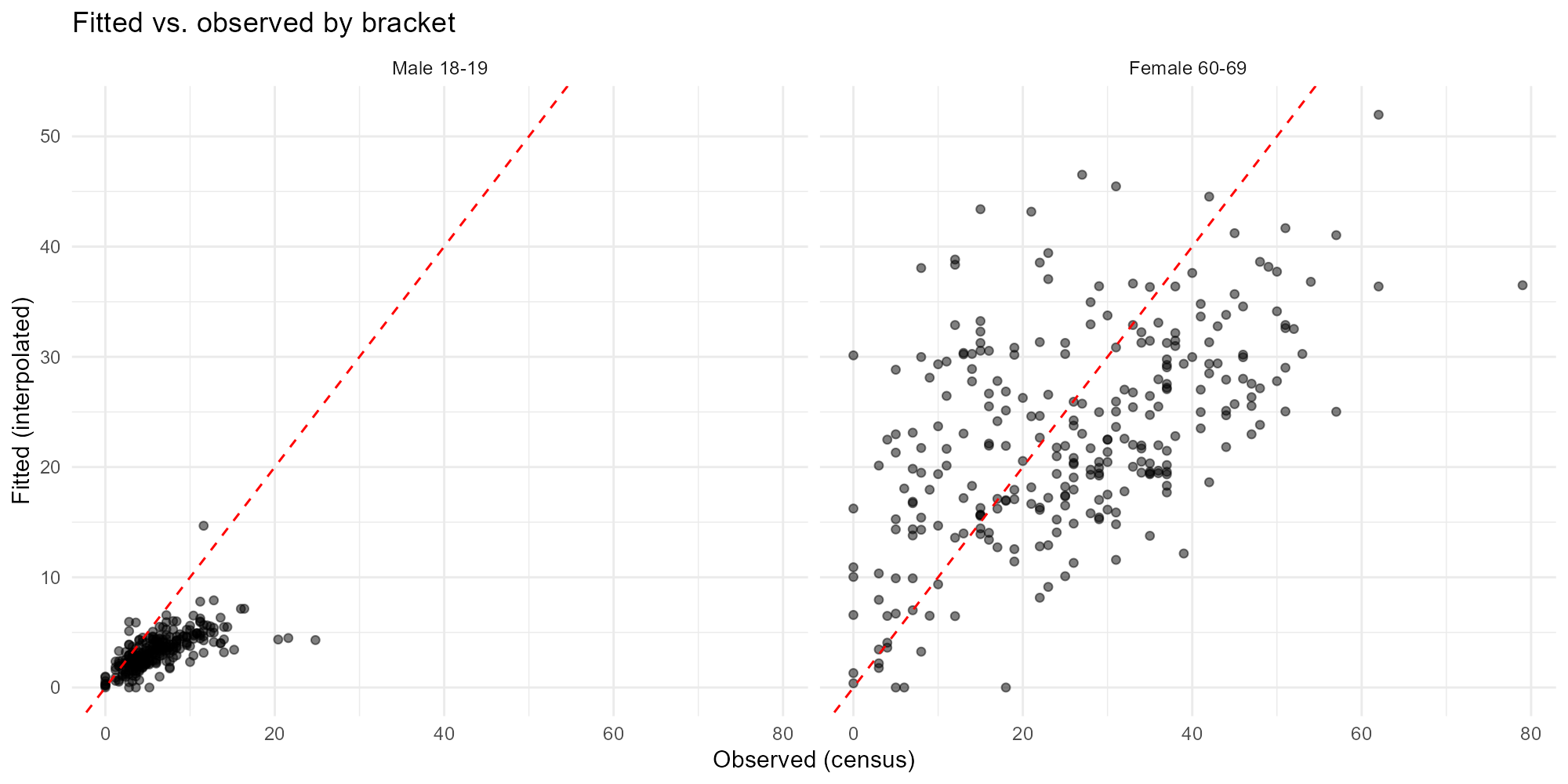
Tabular summary: residual_summary()
residual_summary(result_vga)#> $per_bracket
#> bracket mean rmse max_abs pct_gt_2sd
#> 1 Male 18-19 -2.827957 4.090797 20.49560 11.111111
#> 2 Male 20-24 -4.939068 7.863136 30.09403 10.752688
#> 3 Male 25-29 -3.688171 8.143861 50.85045 5.376344
#> 4 Male 30-39 -8.189961 17.197483 81.67102 8.243728
#> 5 Male 40-49 -6.788528 12.646651 51.29978 7.885305
#> 6 Male 50-59 -3.684587 10.722865 47.07844 5.734767
#> 7 Male 60-69 -2.093190 10.621665 34.50467 3.942652
#> 8 Female 18-19 -1.896057 3.223661 15.23459 8.243728
#> 9 Female 20-24 -4.594982 8.286037 40.84304 6.451613
#> 10 Female 25-29 -3.767024 9.127714 58.36239 4.659498
#> 11 Female 30-39 -8.634406 18.520190 76.77784 9.318996
#> 12 Female 40-49 -7.537632 13.327423 49.34589 10.035842
#> 13 Female 50-59 -4.498207 11.115594 35.52768 7.526882
#> 14 Female 60-69 -2.953405 12.776099 42.50458 3.584229
#>
#> $per_tract
#> tract_id mean rmse worst_bracket
#> 1 317070105000001 -4.46978183 7.780279 Male 30-39
#> 2 317070105000008 -5.69532052 6.521208 Female 30-39
#> 3 317070105000009 -6.40664761 9.267085 Female 50-59
#> 4 317070105000010 -4.09593649 8.241522 Female 60-69
#> 5 317070105000013 -5.79583025 7.723394 Male 50-59
#> 6 317070105000014 -4.70288513 6.429955 Female 60-69
#> 7 317070105000015 -3.21996417 5.024781 Female 20-24
#> 8 317070105000016 -3.66598534 5.970083 Male 40-49
#> 9 317070105000017 -8.02832181 10.453555 Female 30-39
#> 10 317070105000019 -3.39119823 5.524819 Male 60-69
#> 11 317070105000021 -5.09111145 7.203794 Male 40-49
#> 12 317070105000022 -2.92625683 5.141192 Female 25-29
#> 13 317070105000023 -4.70994909 7.473714 Male 30-39
#> 14 317070105000024 -4.65966836 7.084826 Male 25-29
#> 15 317070105000025 -3.58820938 7.515399 Female 60-69
#> 16 317070105000026 -2.59066838 4.599541 Female 30-39
#> 17 317070105000028 -7.85698397 15.446466 Female 30-39
#> 18 317070105000029 -5.40849541 8.450547 Female 40-49
#> 19 317070105000031 -3.89397077 5.834789 Female 60-69
#> 20 317070105000032 -5.28109306 8.720115 Male 20-24
#> 21 317070105000033 -3.80484335 5.778048 Male 60-69
#> 22 317070105000035 -4.22505171 6.261096 Female 30-39
#> 23 317070105000036 -5.25458948 9.366690 Male 30-39
#> 24 317070105000037 -3.69797455 5.705231 Female 60-69
#> 25 317070105000039 -5.18900581 7.768118 Male 40-49
#> 26 317070105000040 -5.54970207 17.902995 Male 50-59
#> 27 317070105000041 -4.35580469 8.416798 Female 50-59
#> 28 317070105000043 -2.92924034 5.086059 Male 40-49
#> 29 317070105000044 -4.92242984 14.221143 Male 40-49
#> 30 317070105000045 -5.54451610 9.378096 Female 40-49
#> 31 317070105000047 -6.87290218 11.932898 Female 50-59
#> 32 317070105000048 -4.12777463 7.548253 Male 30-39
#> 33 317070105000049 -3.86660142 5.565906 Female 30-39
#> 34 317070105000050 -5.07588214 8.838433 Female 60-69
#> 35 317070105000051 -3.16488998 4.579301 Female 40-49
#> 36 317070105000052 -4.98188243 7.269529 Female 30-39
#> 37 317070105000054 -4.35382028 8.647110 Female 60-69
#> 38 317070105000055 -4.29991404 11.477892 Female 50-59
#> 39 317070105000056 -2.65074687 5.708509 Female 60-69
#> 40 317070105000057 -3.53921237 7.376684 Female 60-69
#> 41 317070105000059 -5.54329305 8.078800 Female 40-49
#> 42 317070105000060 -5.67776673 7.052883 Male 50-59
#> 43 317070105000061 -4.54991476 6.796096 Female 50-59
#> 44 317070105000063 -4.27955069 9.015014 Female 60-69
#> 45 317070105000065 -6.46841403 9.618279 Male 60-69
#> 46 317070105000066 -5.61921541 11.229709 Male 60-69
#> 47 317070105000067 -3.70455425 7.798223 Female 50-59
#> 48 317070105000069 -7.08697385 9.610891 Male 30-39
#> 49 317070105000070 -5.11650942 8.360278 Female 50-59
#> 50 317070105000071 -5.00843229 7.873160 Female 60-69
#> 51 317070105000074 -4.61650515 7.532968 Female 60-69
#> 52 317070105000077 -3.93589804 9.000784 Female 60-69
#> 53 317070105000078 -5.27064515 8.735191 Female 60-69
#> 54 317070105000079 -5.42014183 8.153125 Female 40-49
#> 55 317070105000082 -5.65931668 8.814523 Male 60-69
#> 56 317070105000083 -4.67848798 8.477325 Female 50-59
#> 57 317070105000086 -5.79974066 9.232188 Male 40-49
#> 58 317070105000087 -5.33495329 8.472898 Female 50-59
#> 59 317070105000089 -7.47620551 10.905327 Female 40-49
#> 60 317070105000091 -5.88057711 9.118924 Female 60-69
#> 61 317070105000094 -6.92574008 9.256367 Female 60-69
#> 62 317070105000095 -7.06021306 9.379119 Male 40-49
#> 63 317070105000096 -6.87335323 10.553908 Female 40-49
#> 64 317070105000100 -9.94300685 15.623448 Female 40-49
#> 65 317070105000101 -4.73015509 8.957741 Male 50-59
#> 66 317070105000102 -9.45775086 14.596251 Female 50-59
#> 67 317070105000103 -3.42464735 6.877452 Female 60-69
#> 68 317070105000104 -5.05947052 10.247530 Female 60-69
#> 69 317070105000105 -6.64221452 10.466741 Female 60-69
#> 70 317070105000106 -4.74402852 7.739311 Female 60-69
#> 71 317070105000107 -5.04264300 9.476863 Female 60-69
#> 72 317070105000108 -4.52430196 11.836652 Female 60-69
#> 73 317070105000112 -10.70524209 19.208083 Female 40-49
#> 74 317070105000114 -7.58292592 12.414566 Female 50-59
#> 75 317070105000115 -7.85118226 12.988053 Male 50-59
#> 76 317070105000116 -5.23164076 10.780131 Male 50-59
#> 77 317070105000117 -5.95415250 9.466325 Female 40-49
#> 78 317070105000120 -4.89691516 14.875840 Female 30-39
#> 79 317070105000121 -1.44764559 5.226840 Male 50-59
#> 80 317070105000124 -9.30533133 12.488768 Female 30-39
#> 81 317070105000125 -7.83185485 9.944685 Female 40-49
#> 82 317070105000126 -4.28045952 6.748654 Female 60-69
#> 83 317070105000127 -4.77031584 7.512045 Female 50-59
#> 84 317070105000128 -6.51097071 10.078242 Male 60-69
#> 85 317070105000129 -5.90250621 9.063544 Female 50-59
#> 86 317070105000130 -5.64954139 6.886282 Female 50-59
#> 87 317070105000131 -6.75248864 12.220189 Female 60-69
#> 88 317070105000133 -6.03865184 13.555790 Female 60-69
#> 89 317070105000134 -3.08150406 6.888872 Female 50-59
#> 90 317070105000135 -4.89329067 9.373227 Female 60-69
#> 91 317070105000136 -1.84728816 5.654837 Female 30-39
#> 92 317070105000138 -11.01428571 12.405644 Female 60-69
#> 93 317070105000139 -2.83529955 7.100915 Male 50-59
#> 94 317070105000140 -1.73322996 5.343345 Male 50-59
#> 95 317070105000141 -1.96477361 4.927858 Male 50-59
#> 96 317070105000142 -10.51428571 12.016417 Male 40-49
#> 97 317070105000143 -10.95714286 12.349320 Female 40-49
#> 98 317070105000144 -4.29576738 6.475777 Male 40-49
#> 99 317070105000147 -7.57497684 13.883793 Male 40-49
#> 100 317070105000148 -6.95230788 9.746854 Female 40-49
#> 101 317070105000149 -4.04310841 9.315555 Female 50-59
#> 102 317070105000150 -0.09205360 2.453728 Female 60-69
#> 103 317070105000151 -9.38574168 14.562069 Male 50-59
#> 104 317070105000152 -4.05701272 10.191480 Female 60-69
#> 105 317070105000153 -5.12329326 7.086856 Female 60-69
#> 106 317070105000154 -5.97116573 7.561613 Female 50-59
#> 107 317070105000155 -4.40479715 5.876291 Female 50-59
#> 108 317070105000156 -3.38121653 6.447234 Female 60-69
#> 109 317070105000157 -7.38748886 14.083776 Female 40-49
#> 110 317070105000158 -4.51901512 6.005070 Male 60-69
#> 111 317070105000159 -6.55120159 15.163124 Female 40-49
#> 112 317070105000160 -5.82683227 10.990449 Female 50-59
#> 113 317070105000161 -8.37639883 13.662417 Female 50-59
#> 114 317070105000162 -6.40080059 13.818299 Male 50-59
#> 115 317070105000164 -2.37118808 8.546970 Female 30-39
#> 116 317070105000165 0.22212169 1.101596 Female 25-29
#> 117 317070105000166 -6.87200698 9.183925 Female 50-59
#> 118 317070105000168 -7.55816260 9.901026 Female 40-49
#> 119 317070105000169 -6.18064683 8.358678 Female 60-69
#> 120 317070105000171 -5.65975764 14.035889 Female 40-49
#> 121 317070105000172 -5.31781160 16.142278 Male 40-49
#> 122 317070105000174 -4.72435992 16.356485 Male 30-39
#> 123 317070105000175 -2.71496952 10.298496 Female 30-39
#> 124 317070105000176 -1.55969492 3.617670 Female 30-39
#> 125 317070105000177 9.37303532 14.054435 Female 30-39
#> 126 317070105000178 -4.56211561 14.353457 Female 30-39
#> 127 317070105000179 -4.06906832 6.673661 Female 60-69
#> 128 317070105000180 -3.81645096 6.853486 Female 60-69
#> 129 317070105000181 -3.05099036 6.596282 Male 60-69
#> 130 317070105000182 -6.13905913 19.792027 Female 30-39
#> 131 317070105000183 -9.32212146 19.624818 Female 40-49
#> 132 317070105000185 -4.35020716 6.326250 Female 50-59
#> 133 317070105000186 -4.26238139 8.832778 Female 60-69
#> 134 317070105000188 -7.11605725 9.998682 Female 40-49
#> 135 317070105000189 -4.48957226 8.247922 Female 60-69
#> 136 317070105000190 -3.48081790 4.948018 Female 40-49
#> 137 317070105000191 -2.16225685 5.348972 Female 60-69
#> 138 317070105000192 -3.98062714 7.363546 Female 30-39
#> 139 317070105000193 -3.41123408 8.592075 Male 30-39
#> 140 317070105000194 -5.89246770 10.087114 Male 20-24
#> 141 317070105000196 -7.41453811 23.364285 Female 30-39
#> 142 317070105000197 0.08298607 1.586720 Male 30-39
#> 143 317070105000198 -0.52907993 1.760266 Male 60-69
#> 144 317070105000199 -5.66189173 9.579314 Female 40-49
#> 145 317070105000200 -7.39849787 13.993152 Male 40-49
#> 146 317070105000201 -7.37047749 13.272088 Female 50-59
#> 147 317070105000205 -7.30761777 20.104961 Female 40-49
#> 148 317070105000206 -7.38316109 18.697316 Female 40-49
#> 149 317070105000207 -2.19033493 6.801264 Male 20-24
#> 150 317070105000208 -5.74515367 15.286066 Female 30-39
#> 151 317070105000209 -7.17987136 18.356939 Female 30-39
#> 152 317070105000210 -5.18184771 12.734001 Female 40-49
#> 153 317070105000211 -3.76001088 11.339685 Female 40-49
#> 154 317070105000212 -4.63035739 11.521751 Female 40-49
#> 155 317070105000213 -5.17376113 13.426430 Female 30-39
#> 156 317070105000214 -8.88049176 26.172918 Female 30-39
#> 157 317070105000216 -6.60765474 20.959614 Male 30-39
#> 158 317070105000217 -2.98070111 11.574437 Female 30-39
#> 159 317070105000219 -9.00126981 30.011366 Male 30-39
#> 160 317070105000220 -2.24084214 6.325360 Male 20-24
#> 161 317070105000221 -3.40950336 6.094611 Female 60-69
#> 162 317070105000222 -2.09463879 4.456393 Male 60-69
#> 163 317070105000223 -1.54964403 4.448766 Female 60-69
#> 164 317070105000224 -2.54114135 7.189218 Female 25-29
#> 165 317070105000225 -2.06761302 5.207428 Male 60-69
#> 166 317070105000226 -2.03412647 4.494774 Male 25-29
#> 167 317070105000227 -3.93916279 7.105443 Male 30-39
#> 168 317070105000228 -2.88029353 4.856285 Female 20-24
#> 169 317070105000229 -3.72300315 6.301937 Female 60-69
#> 170 317070105000230 -3.40944562 4.990152 Female 60-69
#> 171 317070105000231 -3.85189135 6.035891 Male 30-39
#> 172 317070105000232 -2.85902328 4.248717 Male 60-69
#> 173 317070105000233 -2.60761506 3.980579 Male 30-39
#> 174 317070105000234 -2.04350410 4.884697 Male 20-24
#> 175 317070105000235 -3.92571869 6.701134 Female 30-39
#> 176 317070105000236 -2.29353158 4.713671 Female 20-24
#> 177 317070105000237 -5.05134358 8.760045 Female 40-49
#> 178 317070105000238 -3.10664216 6.845682 Female 60-69
#> 179 317070105000239 -2.58490714 4.991092 Female 50-59
#> 180 317070105000240 -2.55276300 5.017651 Female 60-69
#> 181 317070105000241 -3.29748551 9.692393 Female 50-59
#> 182 317070105000242 -5.47205505 8.967630 Male 30-39
#> 183 317070105000243 -2.94082344 5.089976 Male 30-39
#> 184 317070105000244 -2.92593525 5.871260 Male 20-24
#> 185 317070105000245 -6.09796927 7.880939 Female 40-49
#> 186 317070105000246 -4.79644059 14.507438 Female 30-39
#> 187 317070105000247 -3.87800652 5.314640 Male 25-29
#> 188 317070105000248 -3.64048516 6.197416 Female 60-69
#> 189 317070105000249 -2.70802333 5.142643 Female 60-69
#> 190 317070105000250 -2.14734239 7.188832 Male 40-49
#> 191 317070105000251 -5.85967447 13.289550 Female 30-39
#> 192 317070105000252 -6.13233465 9.426209 Female 40-49
#> 193 317070105000253 -5.62125579 11.996178 Female 30-39
#> 194 317070105000254 -4.43924613 7.247192 Male 50-59
#> 195 317070105000255 -5.17465209 6.778978 Female 50-59
#> 196 317070105000256 -3.54452564 7.116696 Male 40-49
#> 197 317070105000257 -4.56645349 6.532802 Female 40-49
#> 198 317070105000258 -3.57928864 6.598547 Female 40-49
#> 199 317070105000259 -3.75511963 11.034016 Female 60-69
#> 200 317070105000260 -2.68180853 6.567740 Female 60-69
#> 201 317070105000261 -4.53623565 8.572451 Male 60-69
#> 202 317070105000262 -2.86537006 7.349391 Male 60-69
#> 203 317070105000263 -4.95779910 6.752232 Male 40-49
#> 204 317070105000264 -3.41874363 7.568416 Male 60-69
#> 205 317070105000265 -3.59290264 5.707987 Female 50-59
#> 206 317070105000266 -3.09675628 5.397823 Female 60-69
#> 207 317070105000267 -2.82532021 5.326122 Male 40-49
#> 208 317070105000268 -3.39354091 4.775391 Male 30-39
#> 209 317070105000269 -5.82607350 8.532071 Female 60-69
#> 210 317070105000270 -2.58332027 7.494807 Female 40-49
#> 211 317070105000271 -6.86463086 10.141773 Male 40-49
#> 212 317070105000272 -2.99226218 6.048348 Male 20-24
#> 213 317070105000273 -7.42774649 11.595932 Female 40-49
#> 214 317070105000274 -3.59836205 7.065970 Male 40-49
#> 215 317070105000275 -3.57947639 7.495054 Female 50-59
#> 216 317070105000276 -1.15282145 2.235907 Male 50-59
#> 217 317070105000277 -4.94508815 12.877212 Female 50-59
#> 218 317070105000278 -3.56140529 9.012226 Female 60-69
#> 219 317070105000279 -11.38037062 23.160827 Female 30-39
#> 220 317070105000281 -6.46720990 8.330964 Male 30-39
#> 221 317070105000282 -8.32424305 21.559227 Male 40-49
#> 222 317070105000283 -4.29694270 7.084340 Female 60-69
#> 223 317070105000284 -3.58707074 7.900689 Female 60-69
#> 224 317070105000285 -0.51066122 2.584213 Male 50-59
#> 225 317070105000286 -2.05797467 3.537599 Male 60-69
#> 226 317070105000287 -4.27751720 8.201016 Male 50-59
#> 227 317070105000288 -5.98806256 11.224576 Male 50-59
#> 228 317070105000289 -3.97599679 4.952262 Female 40-49
#> 229 317070105000290 -7.56002009 23.456935 Male 30-39
#> 230 317070105000291 -4.25221956 12.654563 Female 30-39
#> 231 317070105000292 -7.19632528 29.571819 Female 30-39
#> 232 317070105000293 -6.73239594 25.289948 Male 30-39
#> 233 317070105000294 -3.44708939 13.481216 Female 40-49
#> 234 317070105000295 -6.78640747 15.223176 Female 30-39
#> 235 317070105000296 -4.09955191 8.383322 Male 60-69
#> 236 317070105000297 -4.10914492 5.277312 Male 20-24
#> 237 317070105000298 -6.16337522 13.886344 Female 30-39
#> 238 317070105000299 -4.35758986 12.768339 Female 30-39
#> 239 317070105000300 -8.75445183 18.876183 Male 30-39
#> 240 317070105000301 -7.11259054 19.989977 Female 30-39
#> 241 317070105000302 -9.18575187 30.215780 Female 30-39
#> 242 317070105000303 -9.82653476 28.642227 Male 30-39
#> 243 317070105000304 -5.07906810 20.678122 Female 30-39
#> 244 317070105000305 -3.59284370 11.397482 Female 30-39
#> 245 317070105000307 -7.31920961 27.990769 Female 30-39
#> 246 317070105000308 -7.75966438 23.471593 Female 25-29
#> 247 317070105000309 -4.00167392 15.882670 Male 30-39
#> 248 317070105000310 -6.15993693 21.472827 Female 25-29
#> 249 317070105000311 -1.57470078 3.016574 Female 60-69
#> 250 317070105000312 -5.54156791 9.630858 Female 40-49
#> 251 317070105000313 -5.85762555 12.702007 Female 40-49
#> 252 317070105000314 -2.87938883 4.196684 Female 30-39
#> 253 317070105000315 -1.37737761 3.483471 Female 50-59
#> 254 317070105000316 -1.23277201 4.479257 Male 40-49
#> 255 317070105000317 0.01399407 2.139608 Male 50-59
#> 256 317070105000318 -0.63718519 3.038524 Female 50-59
#> 257 317070105000319 -10.38046181 34.833139 Female 25-29
#> 258 317070105000320 -7.07545181 25.984031 Male 30-39
#> 259 317070105000321 -2.49054020 4.428701 Male 20-24
#> 260 317070105000322 0.12236642 1.472273 Male 20-24
#> 261 317070105000323 -2.35604319 4.833344 Female 20-24
#> 262 317070105000324 -4.12526054 7.456818 Female 40-49
#> 263 317070105000325 -1.17784209 5.739841 Female 60-69
#> 264 317070105000326 -0.72381730 3.503868 Female 50-59
#> 265 317070105000327 -1.57529433 3.979720 Female 40-49
#> 266 317070105000328 -1.12288164 3.901462 Female 50-59
#> 267 317070105000329 -4.67508709 7.222797 Male 20-24
#> 268 317070105000330 -5.38153777 8.481056 Male 20-24
#> 269 317070105000331 -2.75902925 3.921396 Female 25-29
#> 270 317070105000332 -2.52359254 4.539937 Male 50-59
#> 271 317070105000333 -8.06454046 16.087327 Male 40-49
#> 272 317070105000334 -4.82366537 8.221776 Female 40-49
#> 273 317070105000335 -4.57115619 16.070901 Male 30-39
#> 274 317070105000336 -6.52853053 23.845661 Female 30-39
#> 275 317070105000337 -0.78879429 2.949064 Male 60-69
#> 276 317070105000338 -5.98369692 17.759237 Female 25-29
#> 277 317070105000339 -7.82263327 14.399701 Female 40-49
#> 278 317070105000340 -3.47486827 5.862351 Male 60-69
#> 279 317070105000342 -3.03012118 6.102017 Male 50-59
#>
#> $overall
#> $overall$rmse
#> [1] 11.33309
#>
#> $overall$mean_bias
#> [1] -4.720941
#>
#> $overall$total_deviance
#> [1] 50734.63Interpretation
- Raw residuals: Scale depends on tract population. A residual of 10 means different things in a tract with 50 people vs 5,000. Use Pearson or deviance for fair comparison.
- Pearson residuals: Standardized — values around are typical. Values are outliers under the Poisson assumption.
- Bracket comparison: If one bracket consistently has larger residuals, the model may have trouble with that demographic group — check whether the census data for that bracket is reliable.
- Scatter plot: Points should cluster tightly around the 45-degree line. Systematic departures indicate bias.
5. Weight Structure
Intuition
The weight matrix (dimensions , tracts sources) is the heart of the interpolation. It determines how voters from each station are allocated to each tract. Understanding the structure of reveals:
- Catchment areas: which stations serve which tracts
- Concentration: does a tract depend on one station or many?
- Fragility: tracts dependent on a single station are sensitive to that station’s data
Formal definition
Since is column-normalized (), the row sums are NOT 1. For concentration measures, we row-normalize: .
- Shannon entropy:
- Effective number of sources: — the equivalent number of equal-weight stations
- Herfindahl-Hirschman Index: — ranges from (perfect equality) to 1 (monopoly by one station)
Tabular summary: weight_summary()
head(weight_summary(result_vga), 15)#> tract_id dominant_source dominant_weight top3_weight
#> 317070105000001 317070105000001 26 0.12332540 0.12762603
#> 317070105000008 317070105000008 26 0.04936524 0.09158035
#> 317070105000009 317070105000009 14 0.02734454 0.05627433
#> 317070105000010 317070105000010 14 0.04499412 0.06566122
#> 317070105000013 317070105000013 14 0.02663732 0.06874888
#> 317070105000014 317070105000014 14 0.03383821 0.05306196
#> 317070105000015 317070105000015 10 0.04758405 0.05321897
#> 317070105000016 317070105000016 10 0.01432121 0.03074702
#> 317070105000017 317070105000017 10 0.02586139 0.05960858
#> 317070105000019 317070105000019 9 0.01163259 0.02450621
#> 317070105000021 317070105000021 9 0.01779759 0.03381609
#> 317070105000022 317070105000022 12 0.01063265 0.02620110
#> 317070105000023 317070105000023 12 0.02707778 0.06592539
#> 317070105000024 317070105000024 15 0.01728353 0.04931914
#> 317070105000025 317070105000025 9 0.05604796 0.06651539
#> entropy effective_n_sources herfindahl
#> 317070105000001 0.2588357 1.295421 0.91311103
#> 317070105000008 2.7259793 15.271361 0.12433633
#> 317070105000009 3.2015688 24.571048 0.05991134
#> 317070105000010 2.4463974 11.546674 0.19727550
#> 317070105000013 3.0079064 20.244971 0.07363158
#> 317070105000014 2.9461482 19.032502 0.09822271
#> 317070105000015 1.2610502 3.529126 0.57872449
#> 317070105000016 3.1924598 24.348245 0.05897037
#> 317070105000017 3.2993687 27.095529 0.05022195
#> 317070105000019 3.3866854 29.567785 0.04226690
#> 317070105000021 3.3794205 29.353755 0.04259577
#> 317070105000022 3.2477526 25.732443 0.05030593
#> 317070105000023 2.9090090 18.338616 0.07530382
#> 317070105000024 3.0605082 21.338398 0.06206085
#> 317070105000025 1.9709899 7.177778 0.33450311
#> mean_travel_time_weighted
#> 317070105000001 0.1695345
#> 317070105000008 2.6686783
#> 317070105000009 4.1091487
#> 317070105000010 1.5647360
#> 317070105000013 2.8161189
#> 317070105000014 2.4147949
#> 317070105000015 0.4490260
#> 317070105000016 2.3227200
#> 317070105000017 5.8311386
#> 317070105000019 4.1377839
#> 317070105000021 6.1142955
#> 317070105000022 3.2461322
#> 317070105000023 3.3840604
#> 317070105000024 3.6533201
#> 317070105000025 1.1208944
#>
#> ... (279 rows total)Entropy map: plot_weights(type = "entropy")
plot_weights(result_vga, type = "entropy")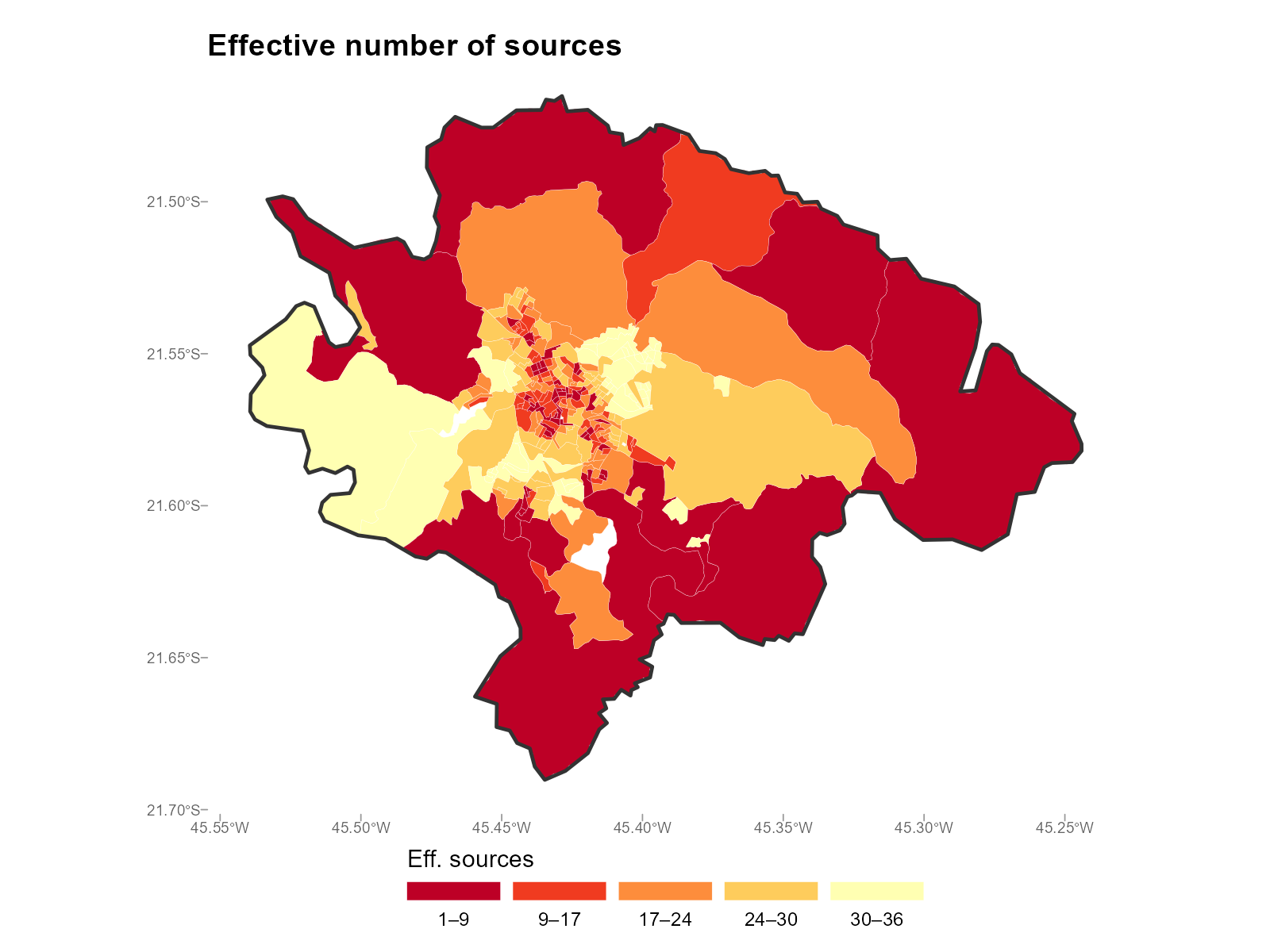
Large city comparison:
plot_weights(result_bh, type = "entropy")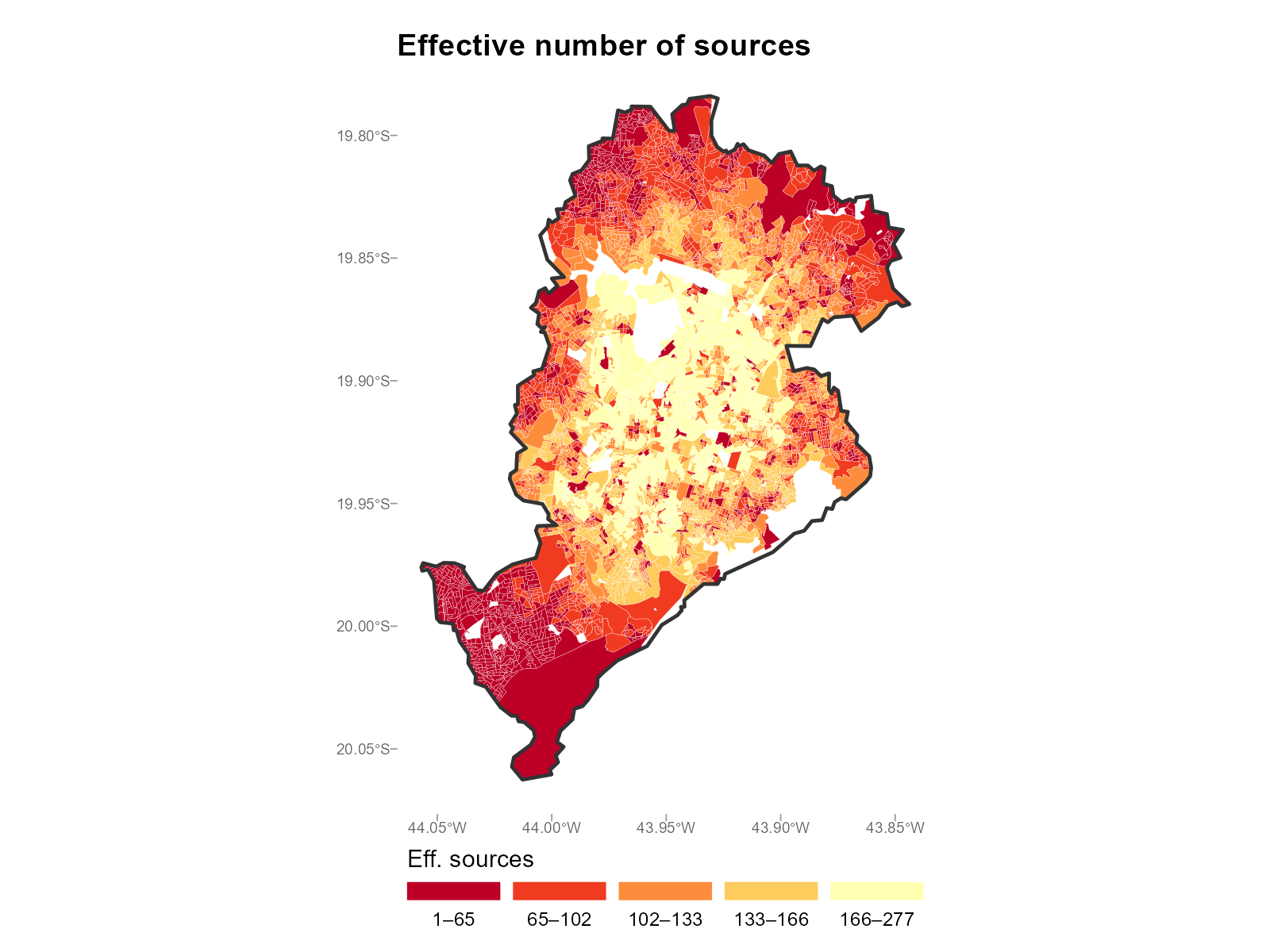
Interactive version (click tracts to see weight summary stats):
plot_weights(result_bh, type = "entropy", interactive = TRUE)Dominant station map:
plot_weights(type = "dominant")
plot_weights(result_vga, type = "dominant")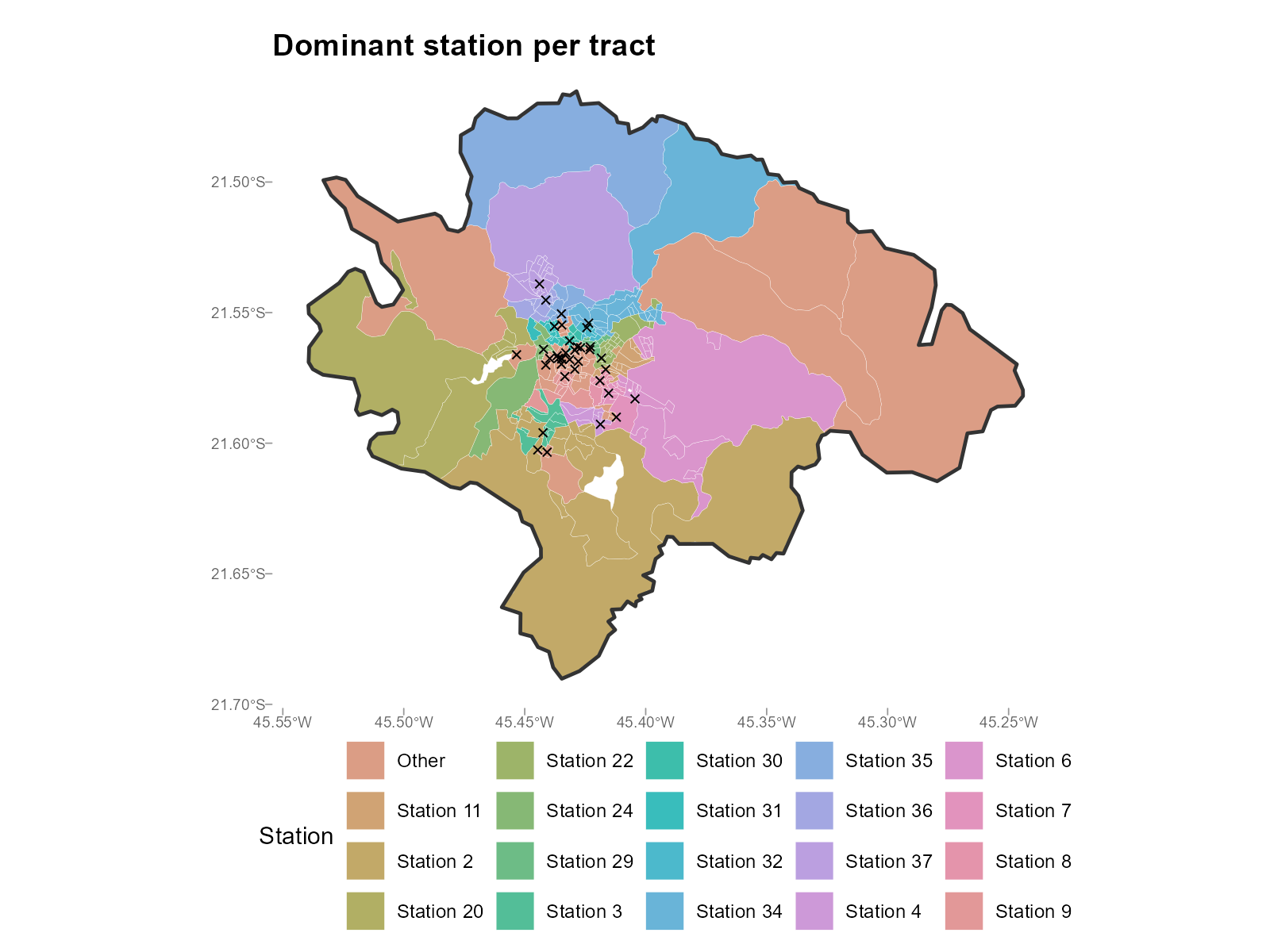
plot_weights(result_nit, type = "dominant")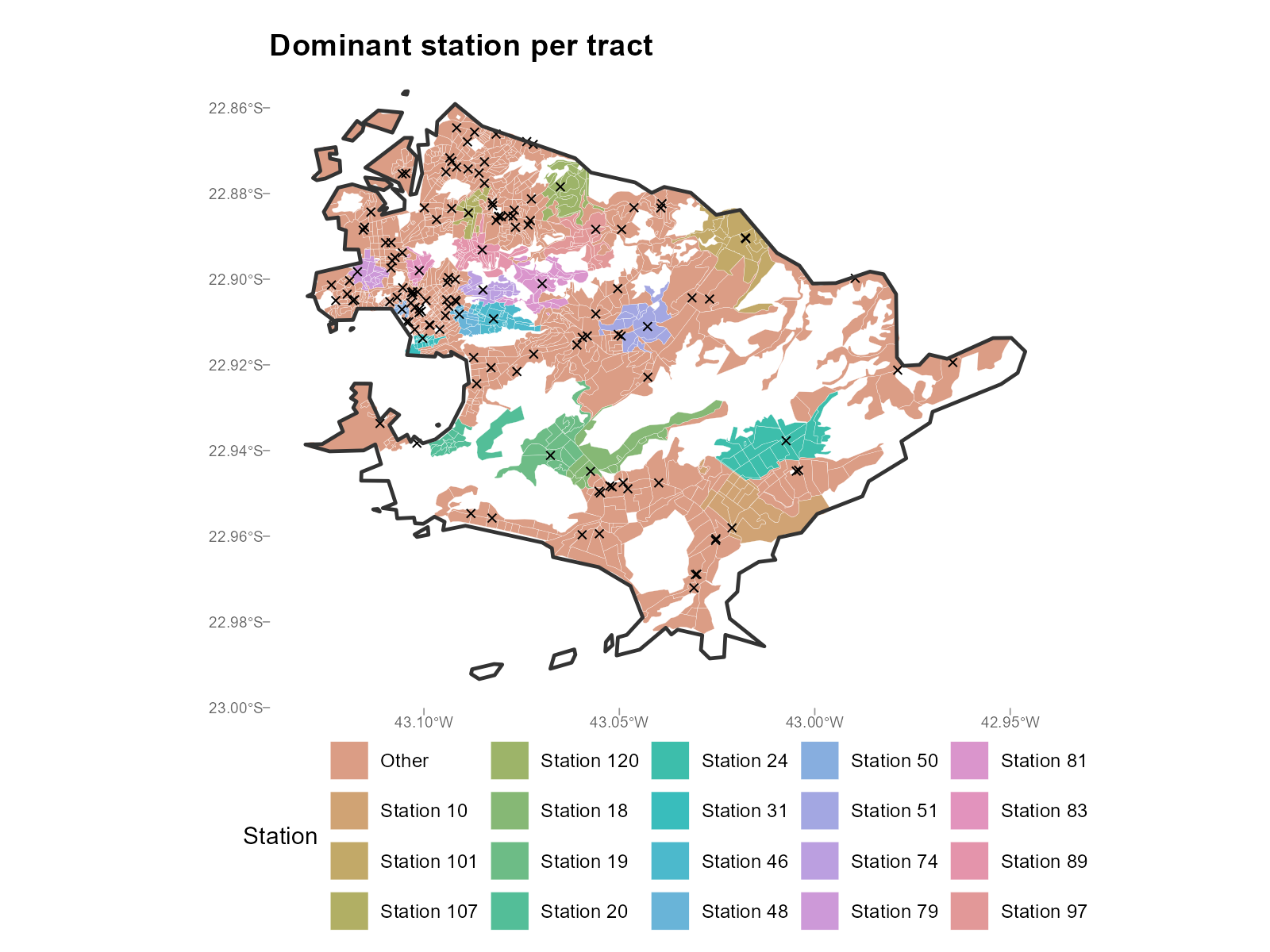
Catchment mode: plot_weights(type = "catchment")
Auto-selects the tract with highest effective number of sources:
plot_weights(result_vga, type = "catchment")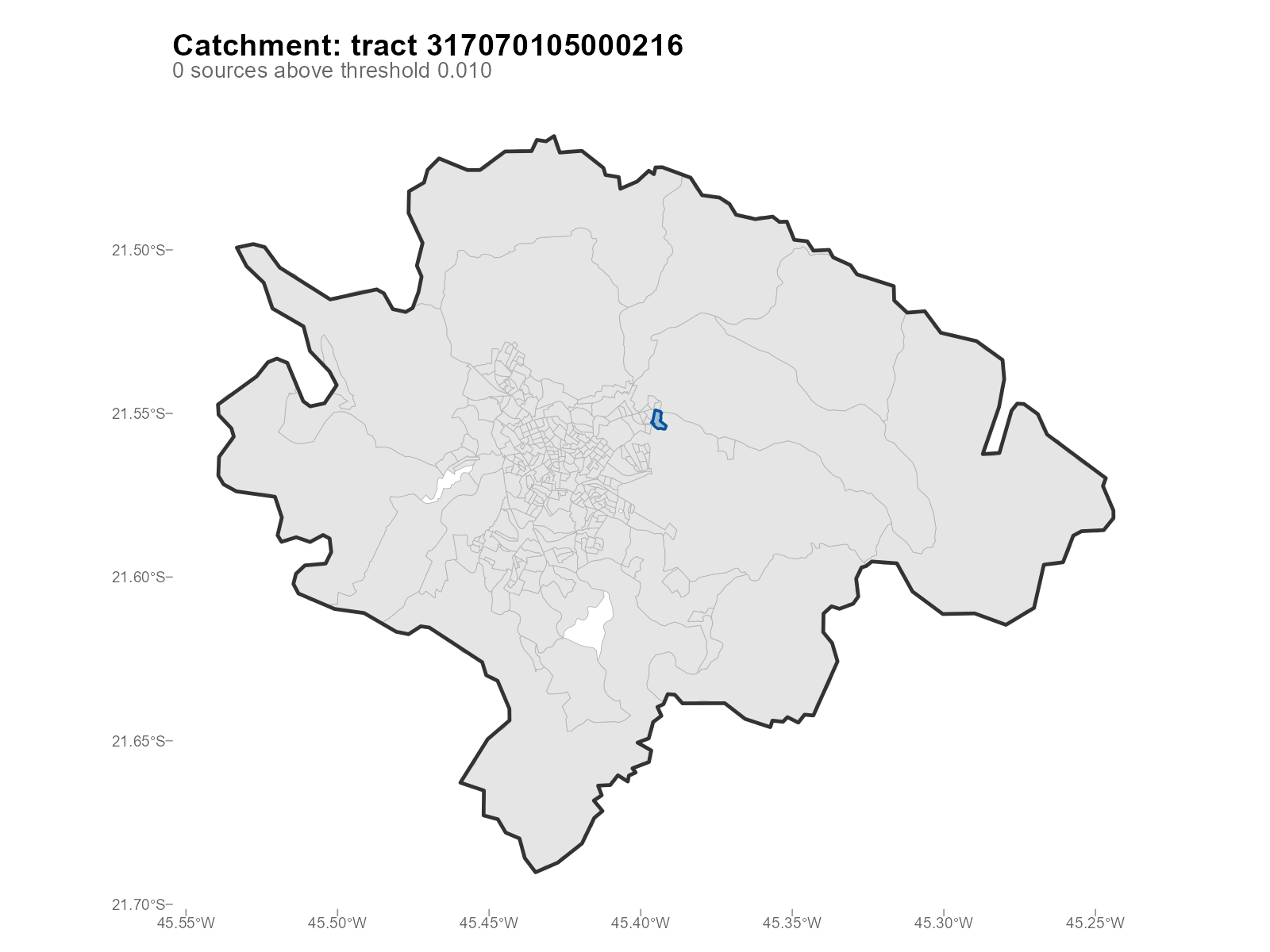
Filtered view with top_k and threshold:
plot_weights(result_vga, type = "catchment", tract = 1,
top_k = 3, threshold = 0.05)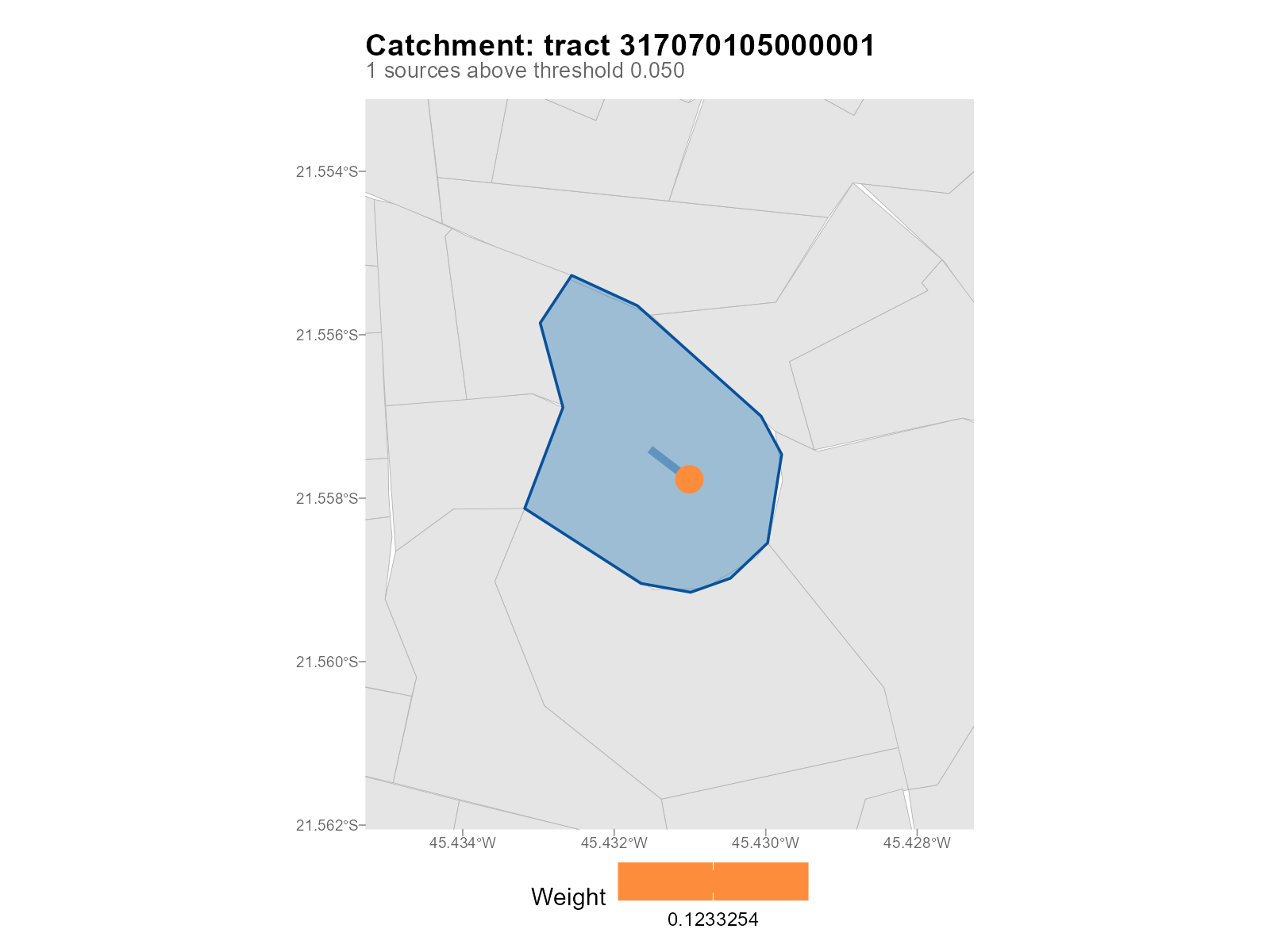
Connections: plot_connections()
Overview — one spoke per tract to its dominant station:
plot_connections(result_vga)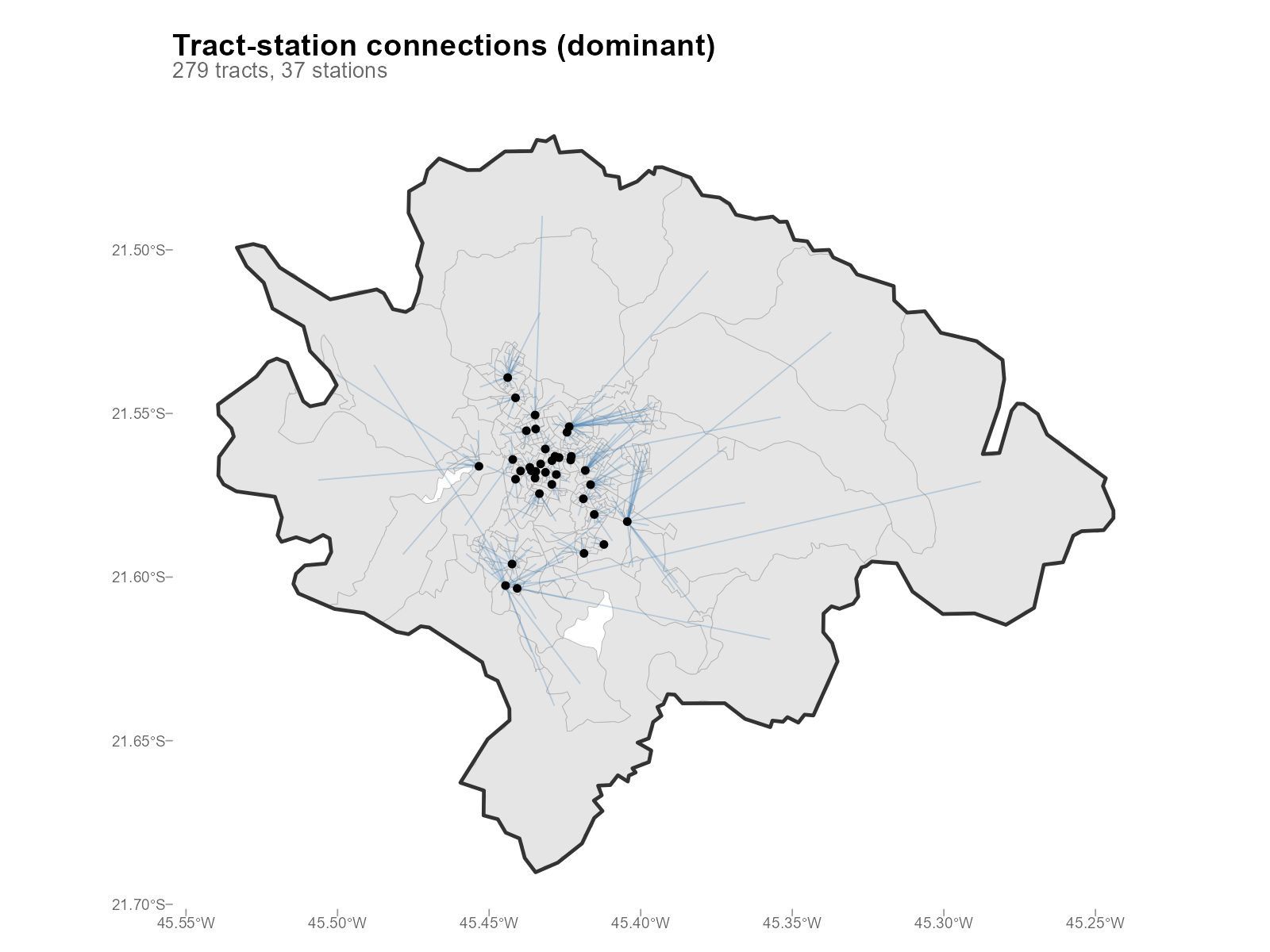
Large city:
plot_connections(result_bh)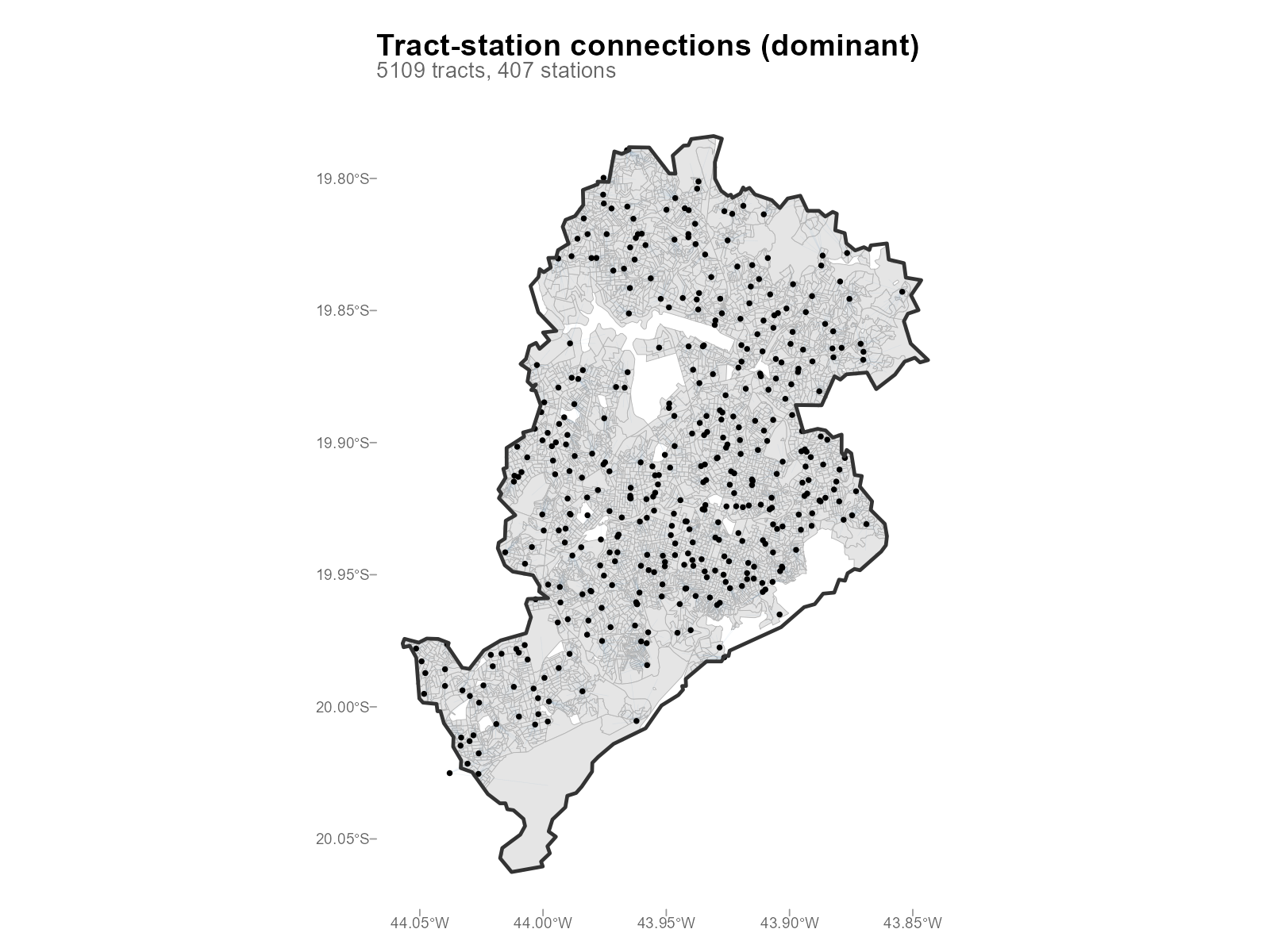
Interactive version for BH (click tracts and stations for detailed weight tables):
Detail for a specific tract:
plot_connections(result_vga, tract = 1)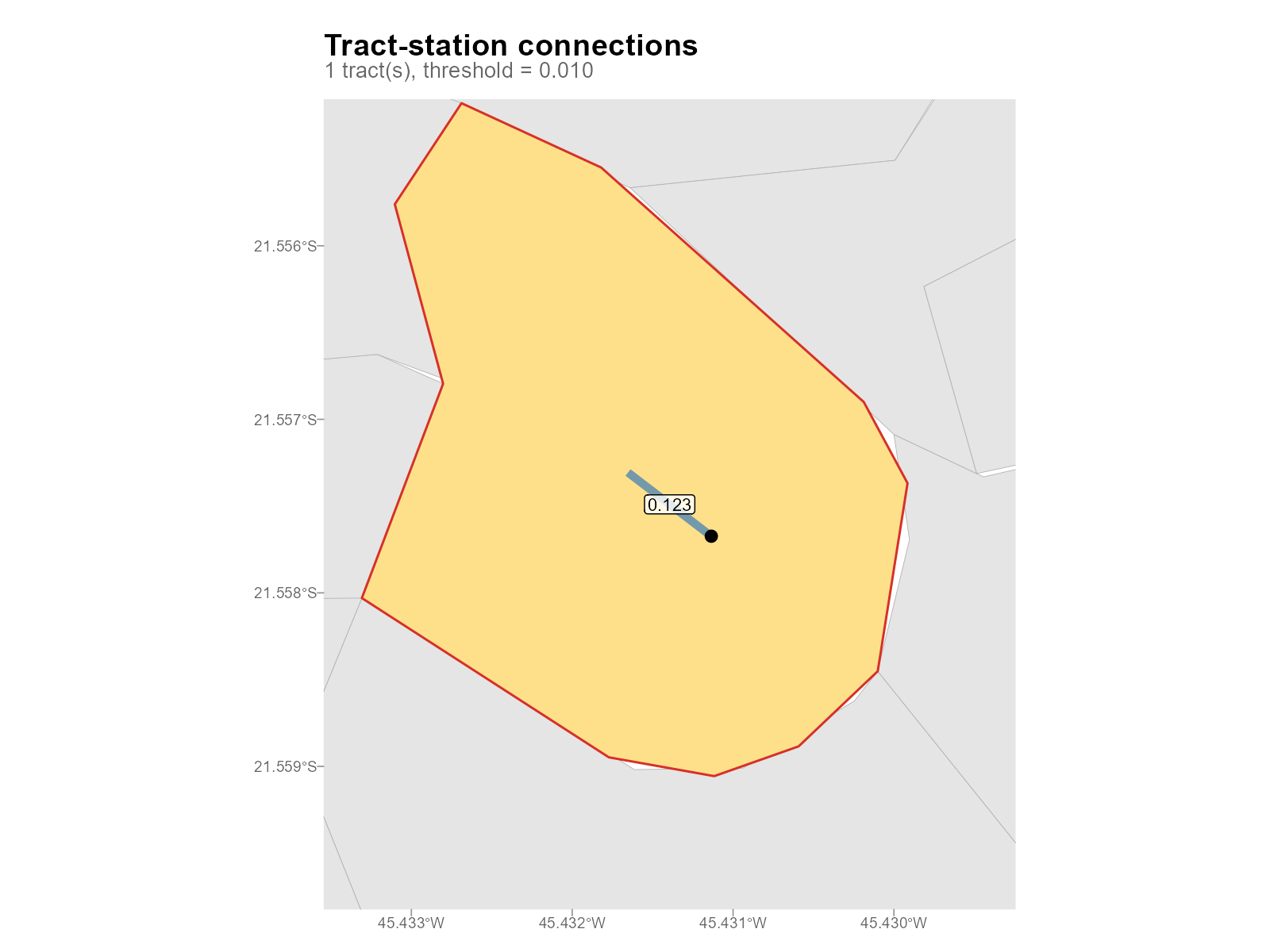
Interactive exploration
All weight diagnostics support interactive = TRUE, which
produces leaflet/mapview maps with rich click popups. This is the
recommended way to explore the weight structure for individual tracts
and stations.
Interactive connections — click any tract to see its top-5 connected stations (with weights, travel times, and cumulative weight). Click any station to see which tracts it serves:
plot_connections(result_vga, interactive = TRUE)Interactive dominant station map — click any tract to see weight summary statistics (effective sources, HHI, dominant weight):
plot_weights(result_vga, type = "dominant", interactive = TRUE)Interpretation
-
Effective number of sources: This is the
exponential of Shannon entropy on the row-normalized weights. A value of
1 means the tract depends on a single station; a value of 10 means the
weights are equivalent to 10 equal-weight stations. High values (e.g.,
> 50) can indicate that distant stations are receiving non-negligible
weight. The default
max_trip_duration = 180(3 hours walking) andfill_missing = NApolicy (see Section 6) address this by assigning exactly zero weight to unreachable and distant pairs. - Low entropy tracts (effective N close to 1): The interpolation for this tract depends almost entirely on a single station. If that station’s data is atypical, the interpolated values may be unreliable.
- Dominant station map: Produces a Voronoi-like partition. If catchment areas don’t match geographic intuition, check the travel time matrix.
- Connections: The spoke pattern should make geographic sense — tracts should connect to nearby stations. Long-distance connections suggest routing issues.
6. Travel Time Data Quality: plot_travel_times()
Intuition
The travel time matrix is the input that drives everything: optimization, weight computation, and ultimately the spatial allocation of voters. Poor travel time data (unreachable pairs, disconnected road networks) directly degrades interpolation quality.
Applied examples
Histogram — distribution of all travel times:
plot_travel_times(result_vga, type = "histogram")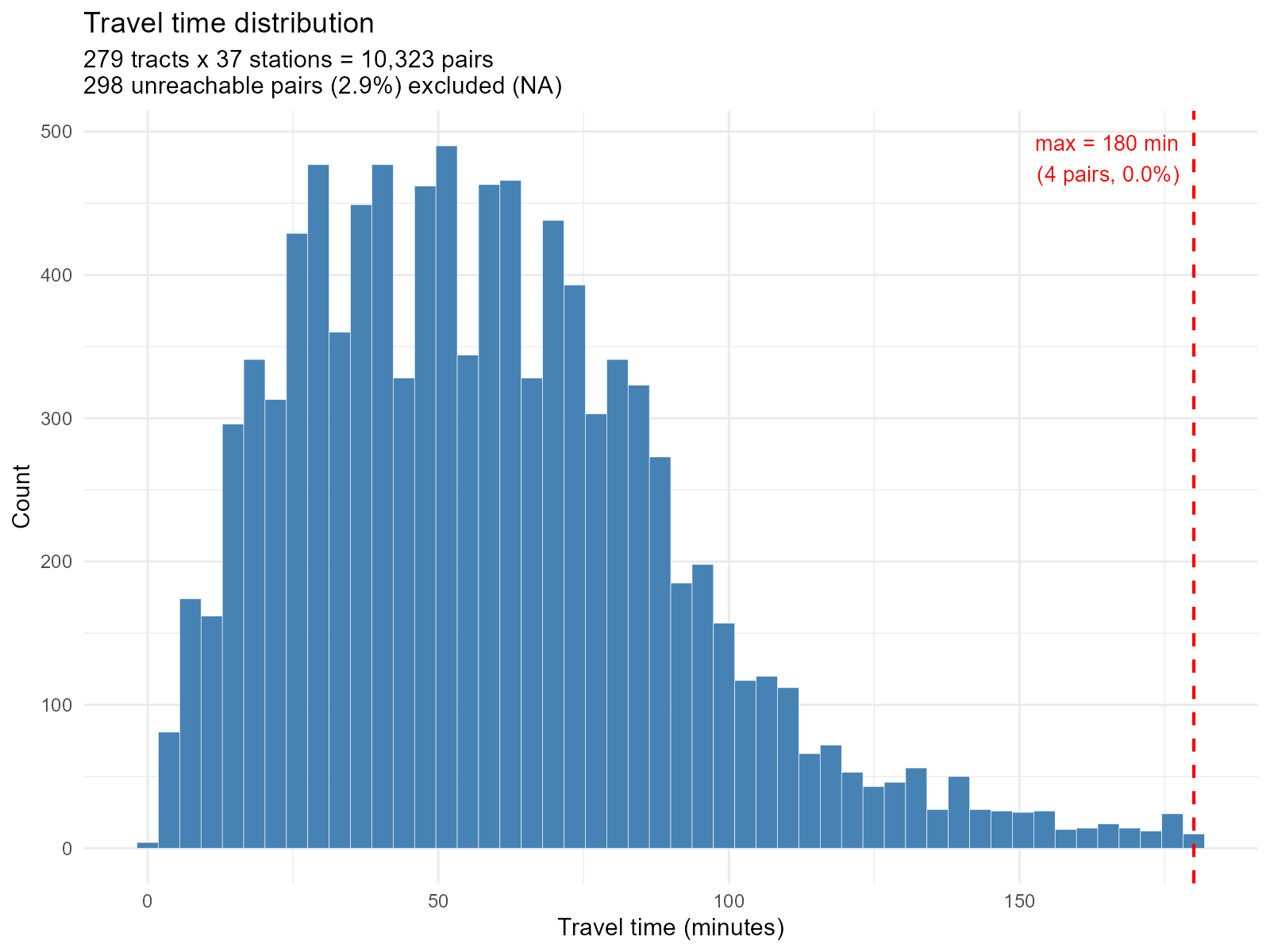
Large city:
plot_travel_times(result_bh, type = "histogram")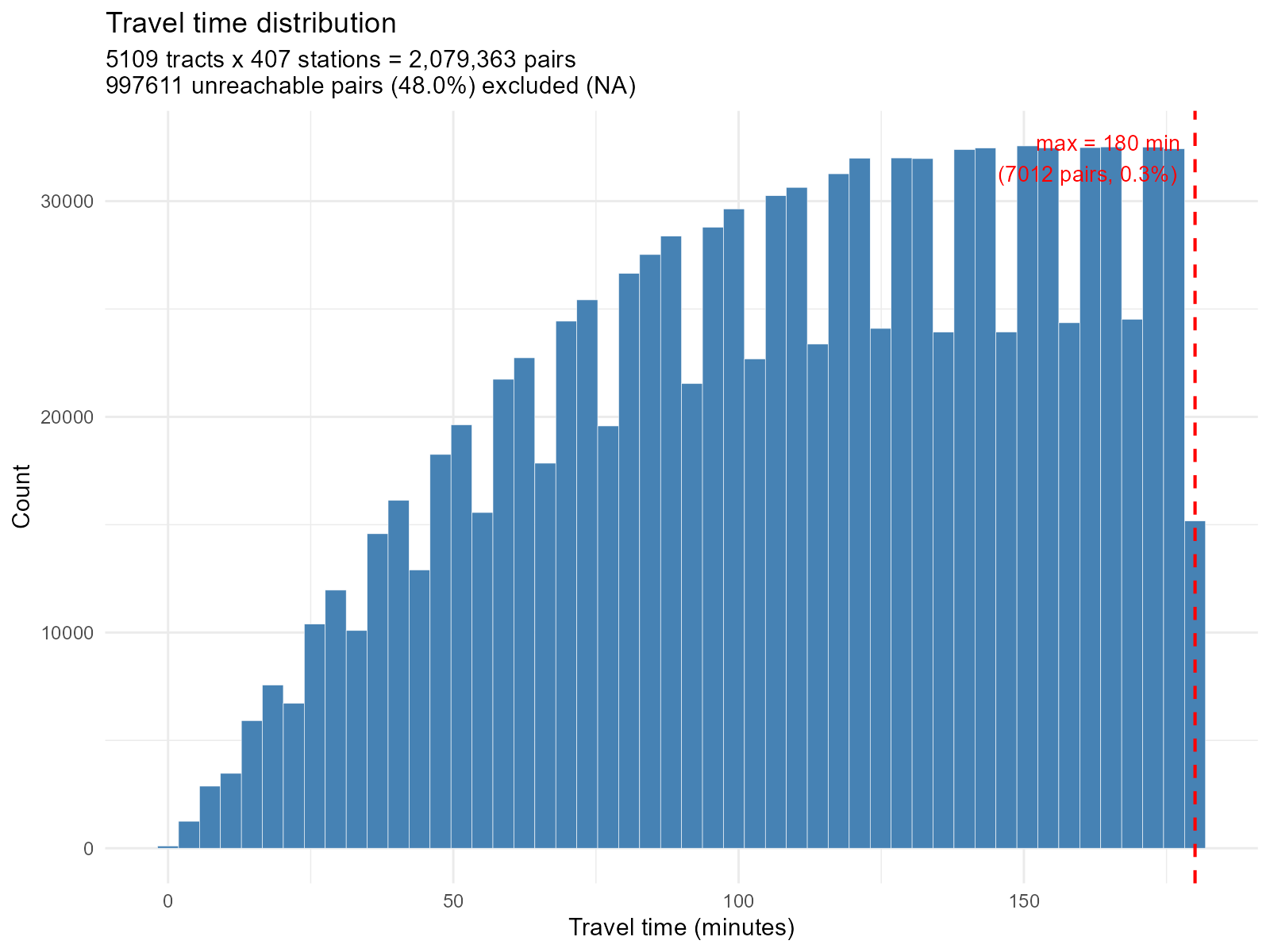
Heatmap — rows = tracts (ordered by latitude), columns = stations. Block-diagonal structure reveals spatial locality:
plot_travel_times(result_vga, type = "heatmap")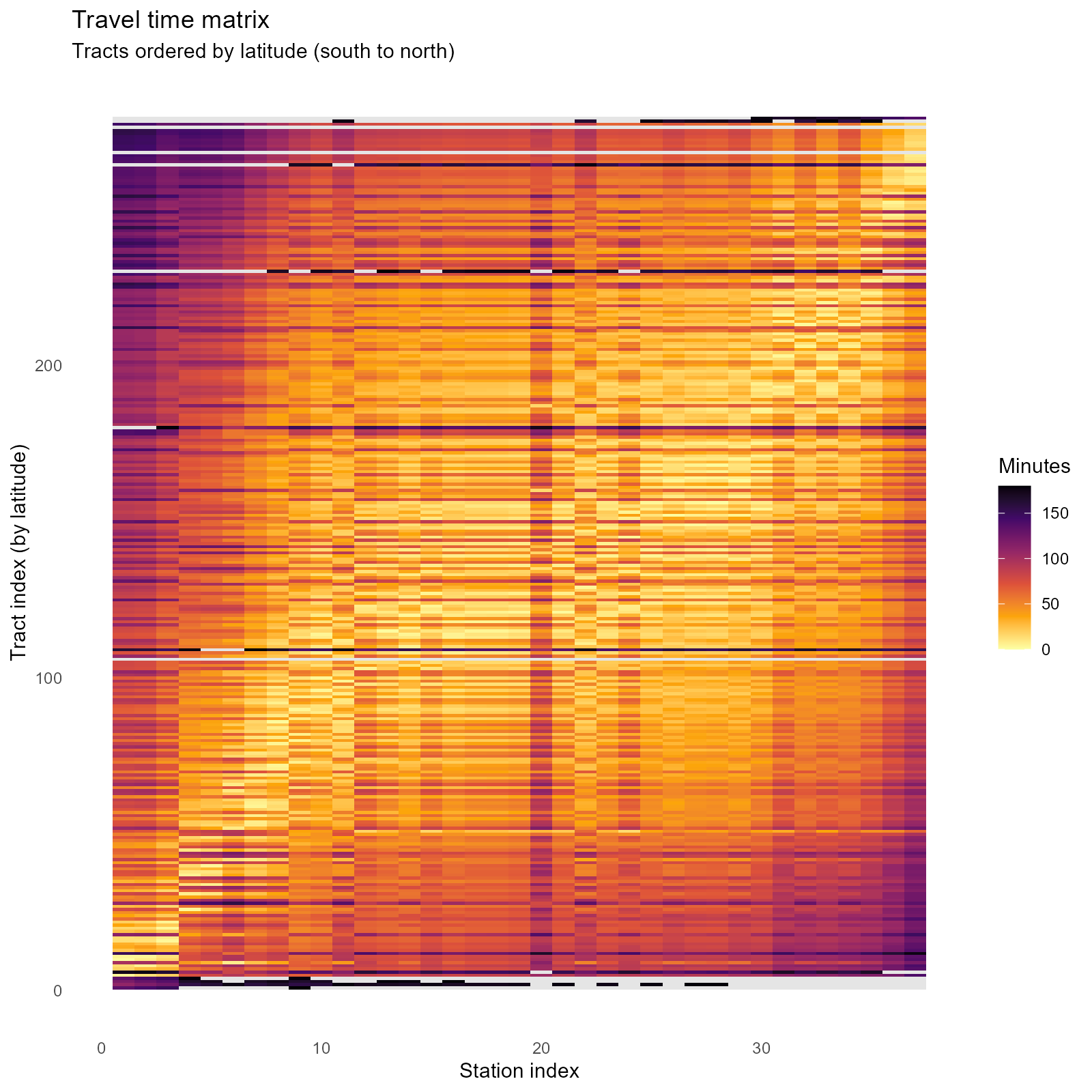
Map — station points colored by travel time for a specific tract:
plot_travel_times(result_vga, type = "map", tract = 1)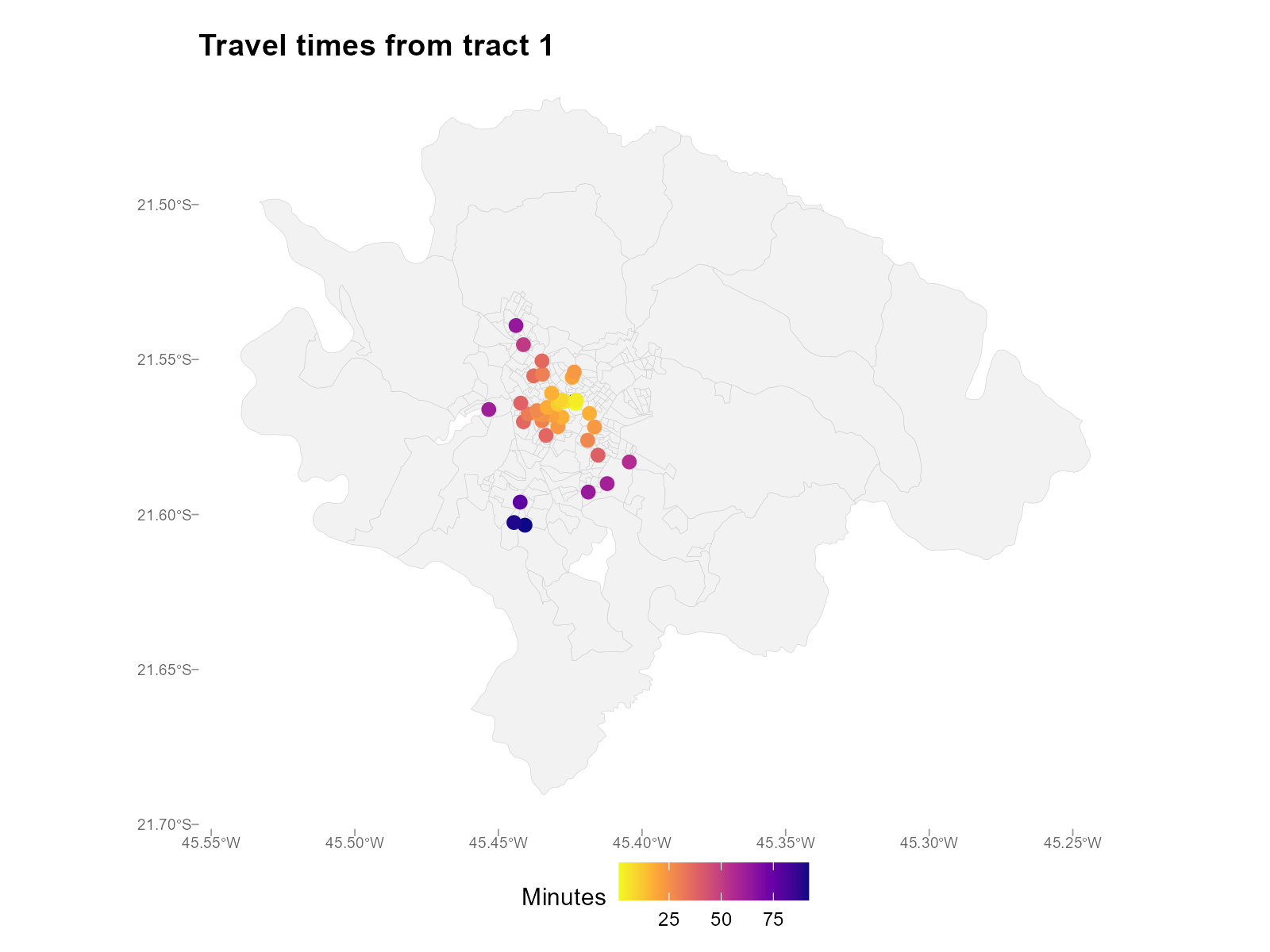
Unreachable pairs and the fill_missing parameter
When the routing engine cannot find a path between a tract and a
station within max_trip_duration minutes, the travel time
is NA. By default,
routing_control(fill_missing = NULL) resolves to
NA, which means these unreachable pairs remain as
NA in the travel time matrix and receive exactly
zero weight — they are completely excluded from both the
optimization and the interpolation.
The default max_trip_duration = 180 (3 hours walking)
intentionally cuts distant links. Any tract–station pair beyond this
threshold is treated as unreachable. This prevents distant stations from
creating a uniform background weight that inflates the effective number
of sources.
If a tract has no reachable stations at all (entire row is
NA), it is excluded from optimization and receives
NA interpolated values.
To assign a finite fill value instead (e.g., to ensure every pair has a travel time):
routing_control(fill_missing = 300) # fill unreachable with 300 minInterpretation
-
NA pairs in histogram: The subtitle reports how
many pairs are unreachable (NA). A large number suggests poor OSM
network coverage or a restrictive
max_trip_duration. -
Grey cells in heatmap: Represent
NA(unreachable) pairs. These receive zero weight. - Bimodal distribution: May indicate disconnected graph components (e.g., separated by a river without bridge in the OSM network).
- Block-diagonal heatmap: Expected — nearby tracts have similar travel times to the same stations. Absence of block structure suggests the travel times lack spatial coherence.
7. Spatial Autocorrelation:
plot_residual_autocorrelation() +
plot_moran()
Intuition
Residual autocorrelation: If residuals are spatially clustered (nearby tracts have similar residuals), the model is missing spatial structure — a misspecification signal.
Moran’s I on interpolated variables: If the interpolated vote shares show expected spatial concentration (e.g., PT strongholds in low-income areas), the method is preserving realistic political geography.
Formal definition
Global Moran’s I:
where are spatial weights (queen contiguity). Range: . Positive = clustering, = random, negative = dispersion.
LISA (Local Moran’s I): . Classification: HH (high-high cluster), LL (low-low), HL (high surrounded by low), LH (low surrounded by high).
Applied examples: Residual autocorrelation
plot_residual_autocorrelation(result_vga)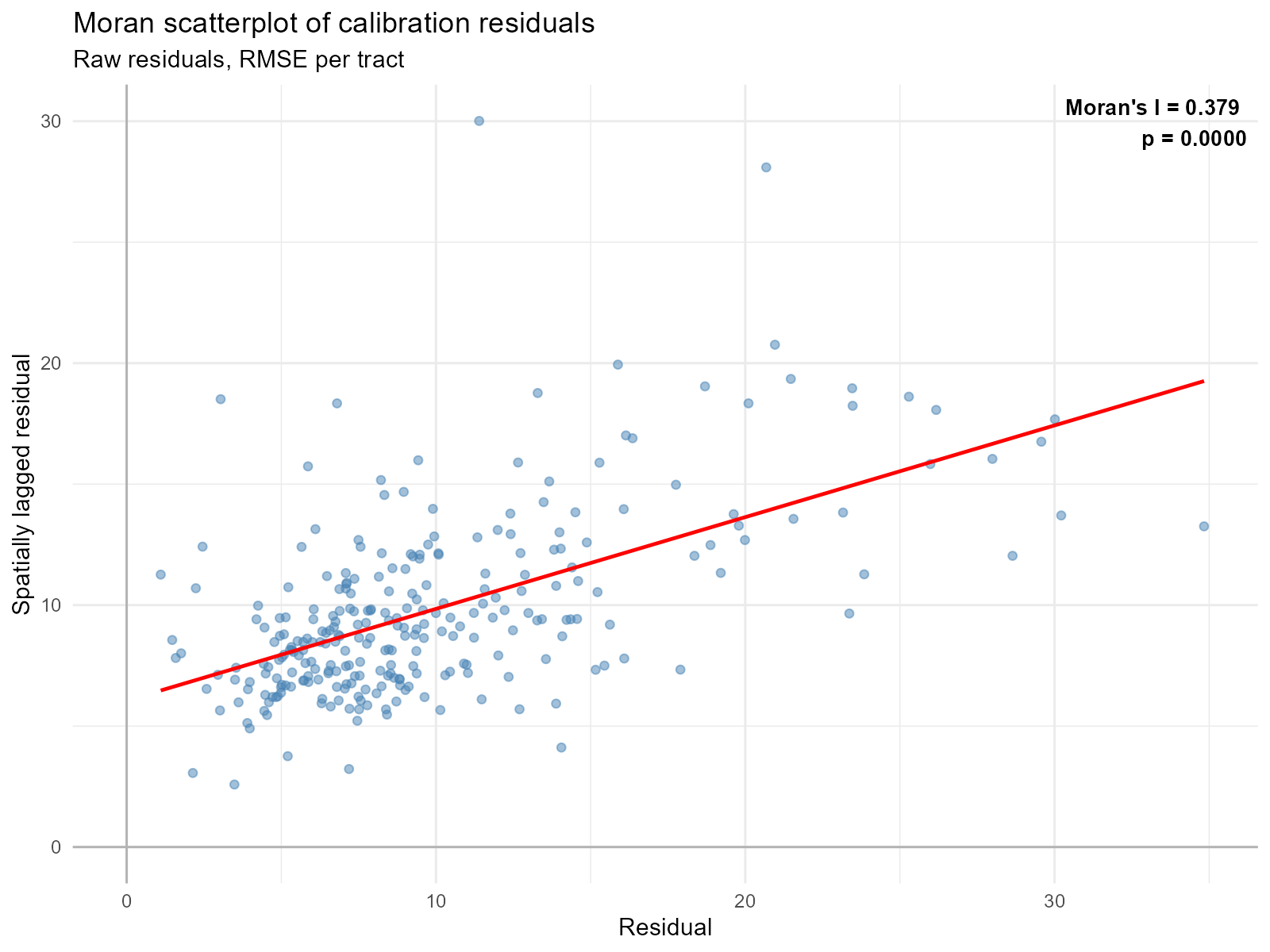
Pearson residuals:
plot_residual_autocorrelation(result_vga, residual_type = "pearson")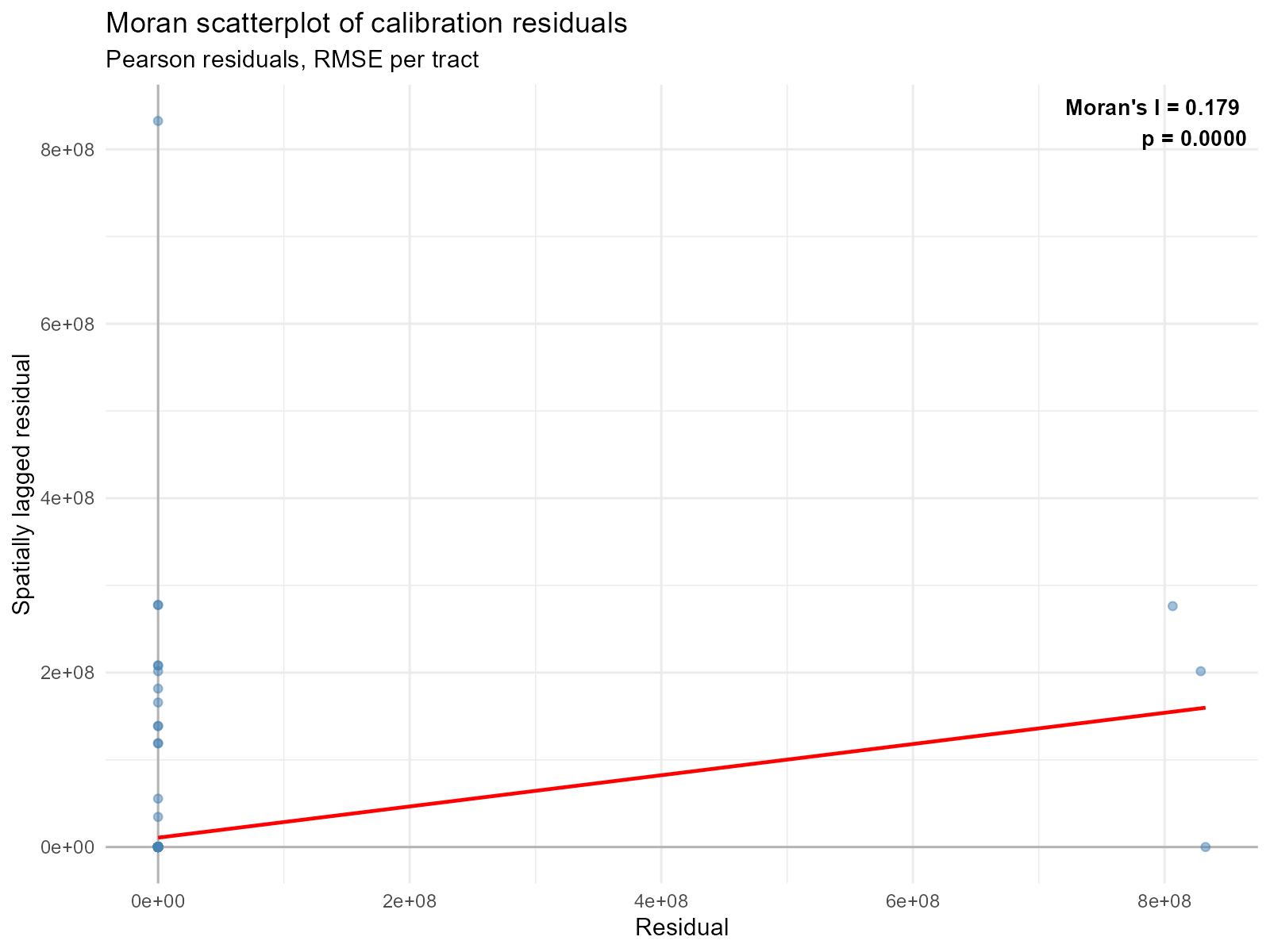
Applied examples: LISA cluster maps
plot_moran(result_vga, variable = "Lula", type = "lisa")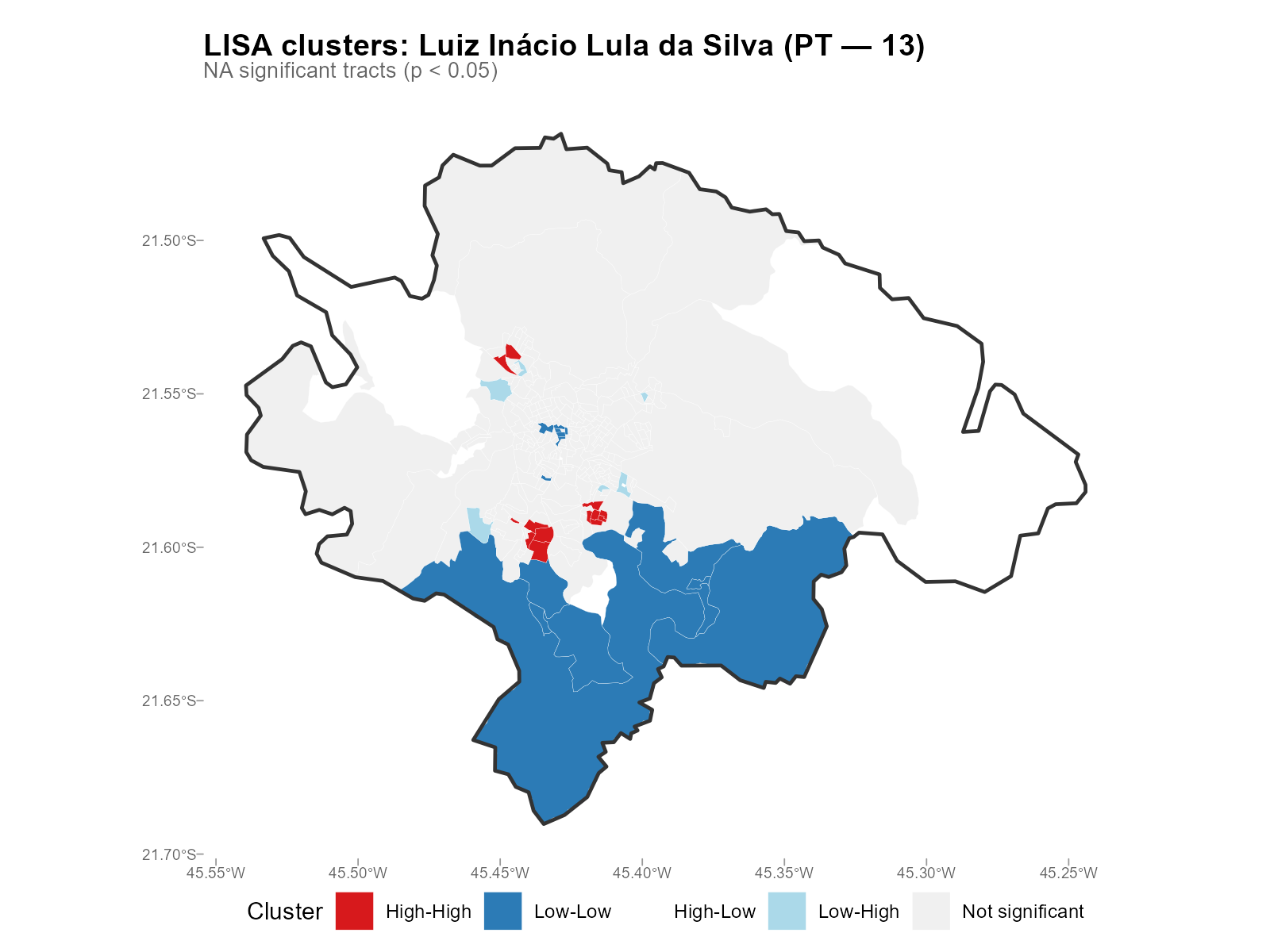
Moran scatterplot:
plot_moran(result_vga, variable = "Lula", type = "moran")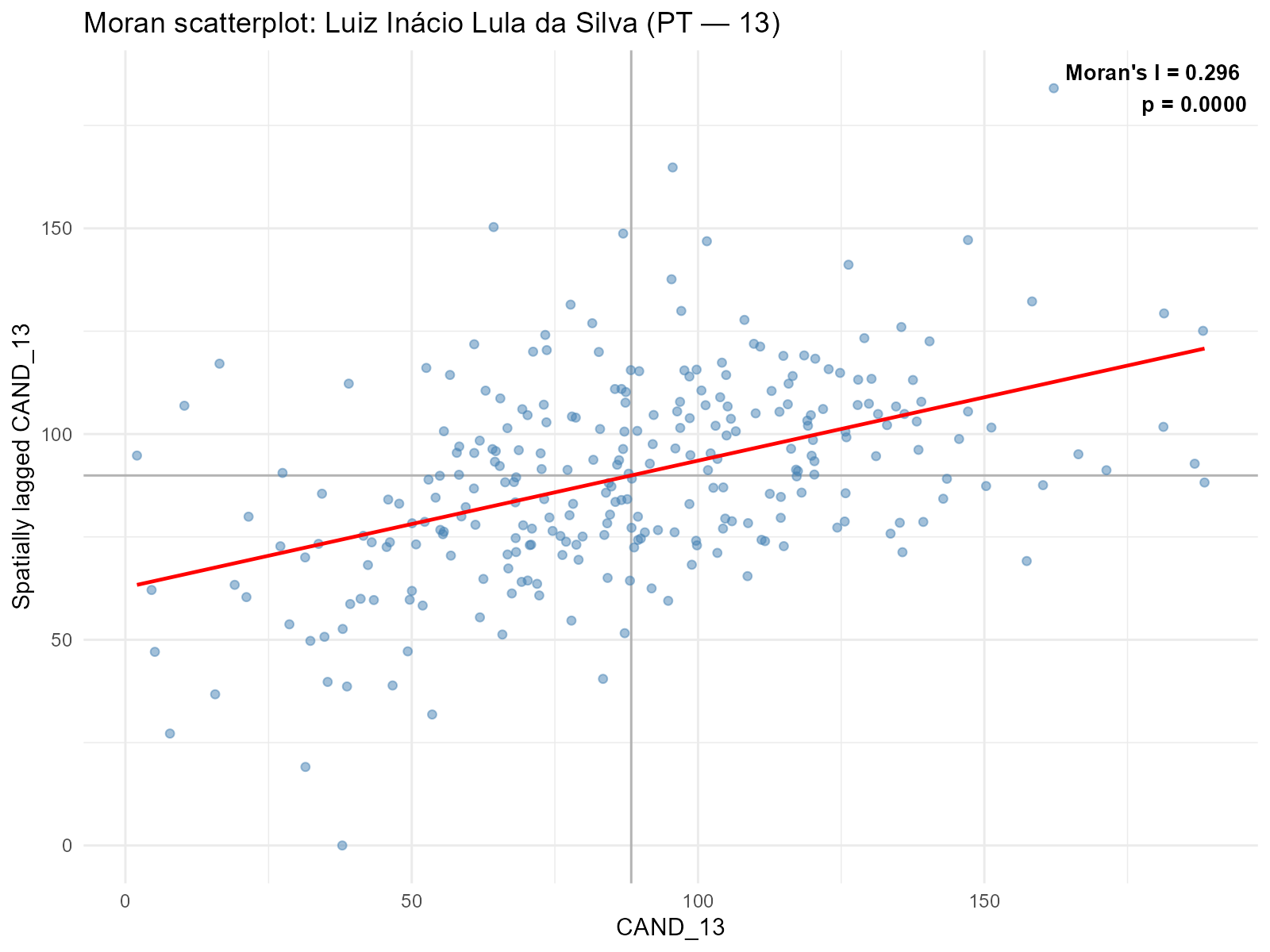
Large city:
plot_moran(result_bh, variable = "Lula", type = "lisa")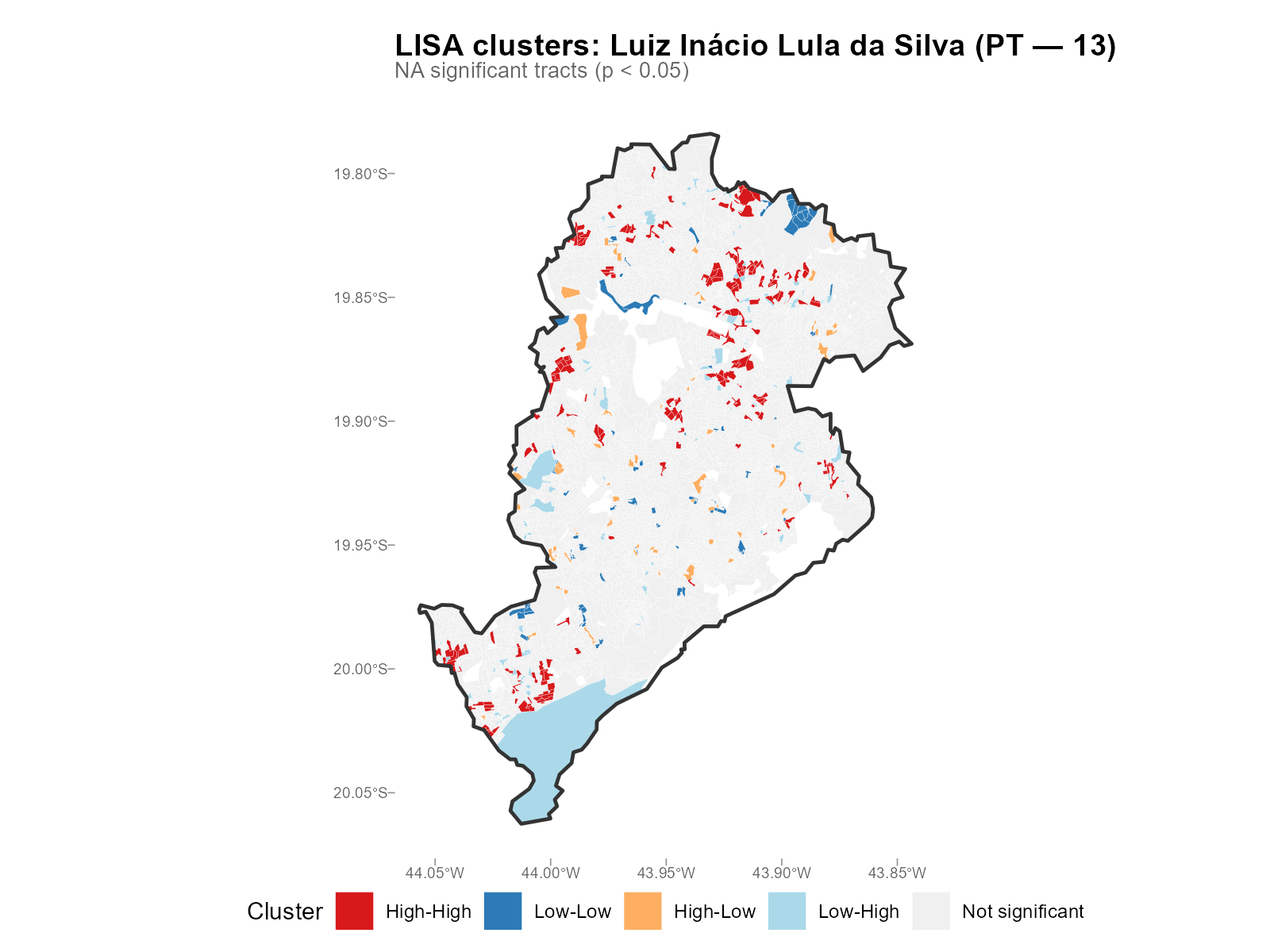
By party:
plot_moran(result_nit, variable = "PT", type = "lisa")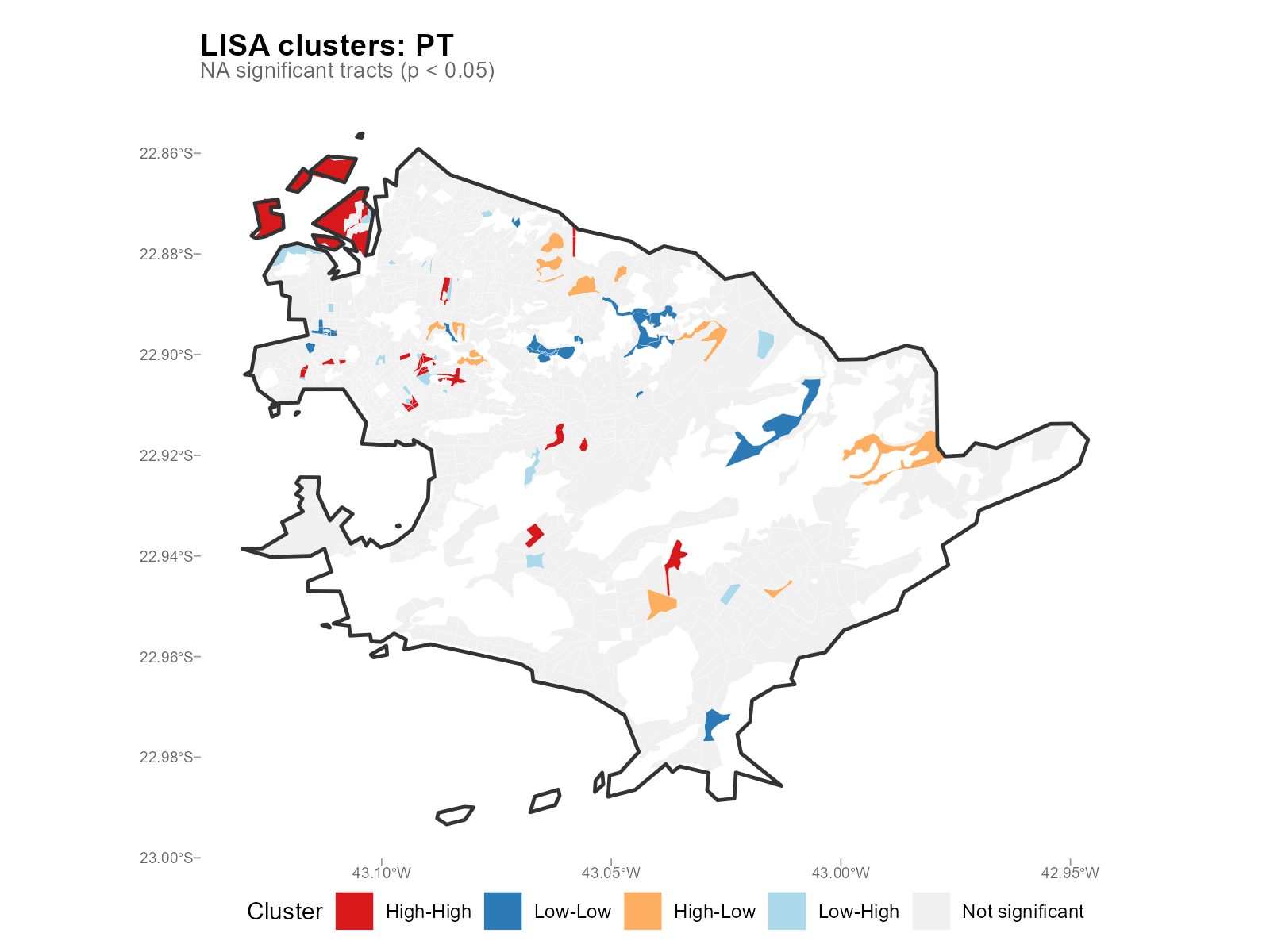
Interpretation
- Significant Moran’s I on residuals (p < 0.05): The model has spatially structured error. Consider whether the travel time data has systematic problems in certain areas.
- HH clusters of Lula votes: Expected in low-income neighborhoods (PT strongholds). Their presence validates that the interpolation preserves known political geography.
- HL/LH clusters: Spatial outliers — tracts with high vote share surrounded by low (or vice versa). May indicate local effects or data irregularities.
8. Ecological Validation
8a. Implied Turnout Rates: plot_turnout_rates()
Intuition
For each tract and demographic bracket, the implied turnout rate is:
In Brazil, voting is compulsory for ages 18–69, so expected turnout is ~75–85%. For 16–17 and 70+, voting is optional and rates should be lower. Rates exceeding 100% indicate tracts where the model allocates more voters than residents — signaling boundary effects or registration mismatches.
Applied examples
Bracket boxplot:
plot_turnout_rates(result_vga, type = "bracket")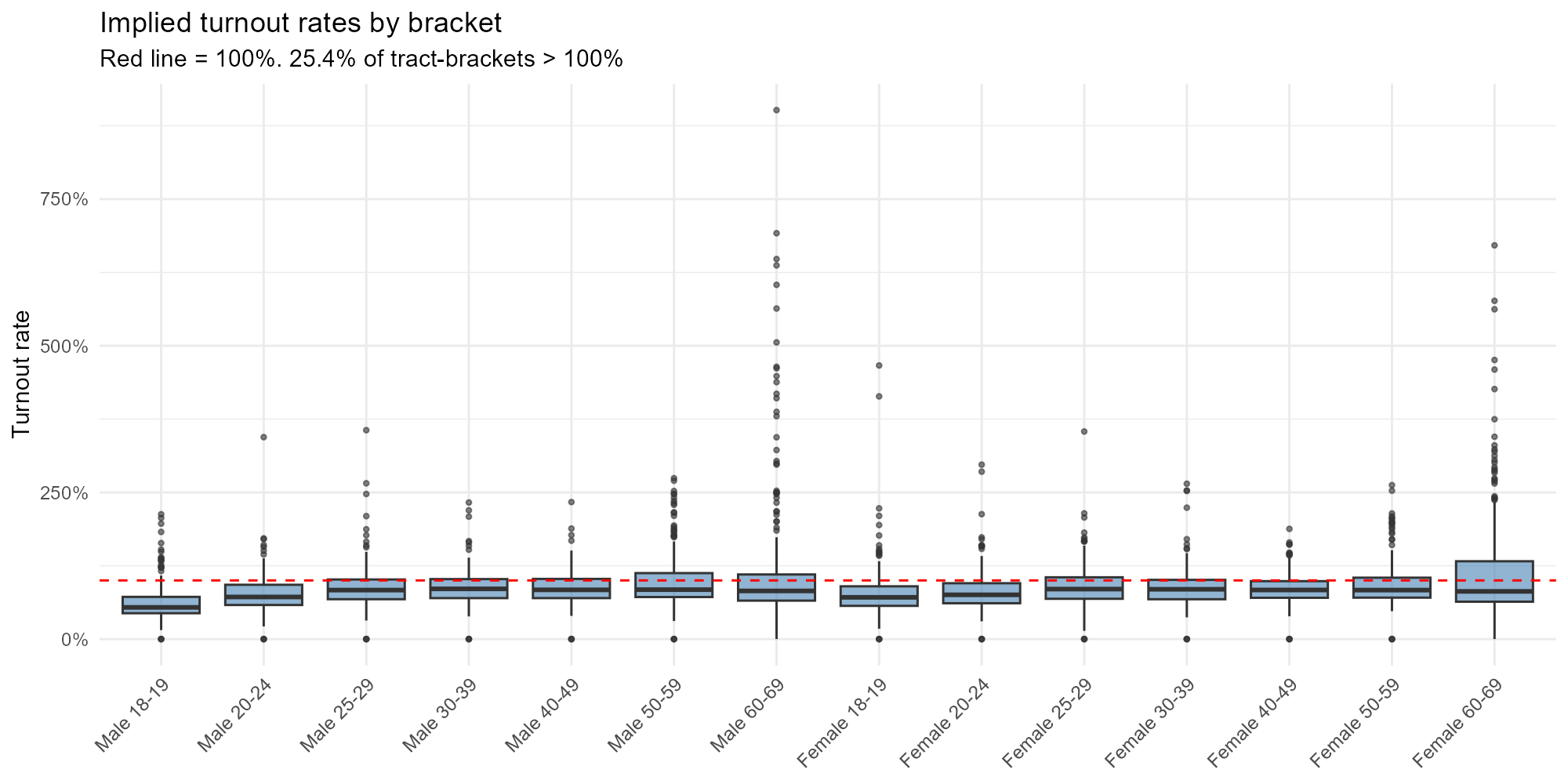
plot_turnout_rates(result_nit, type = "bracket")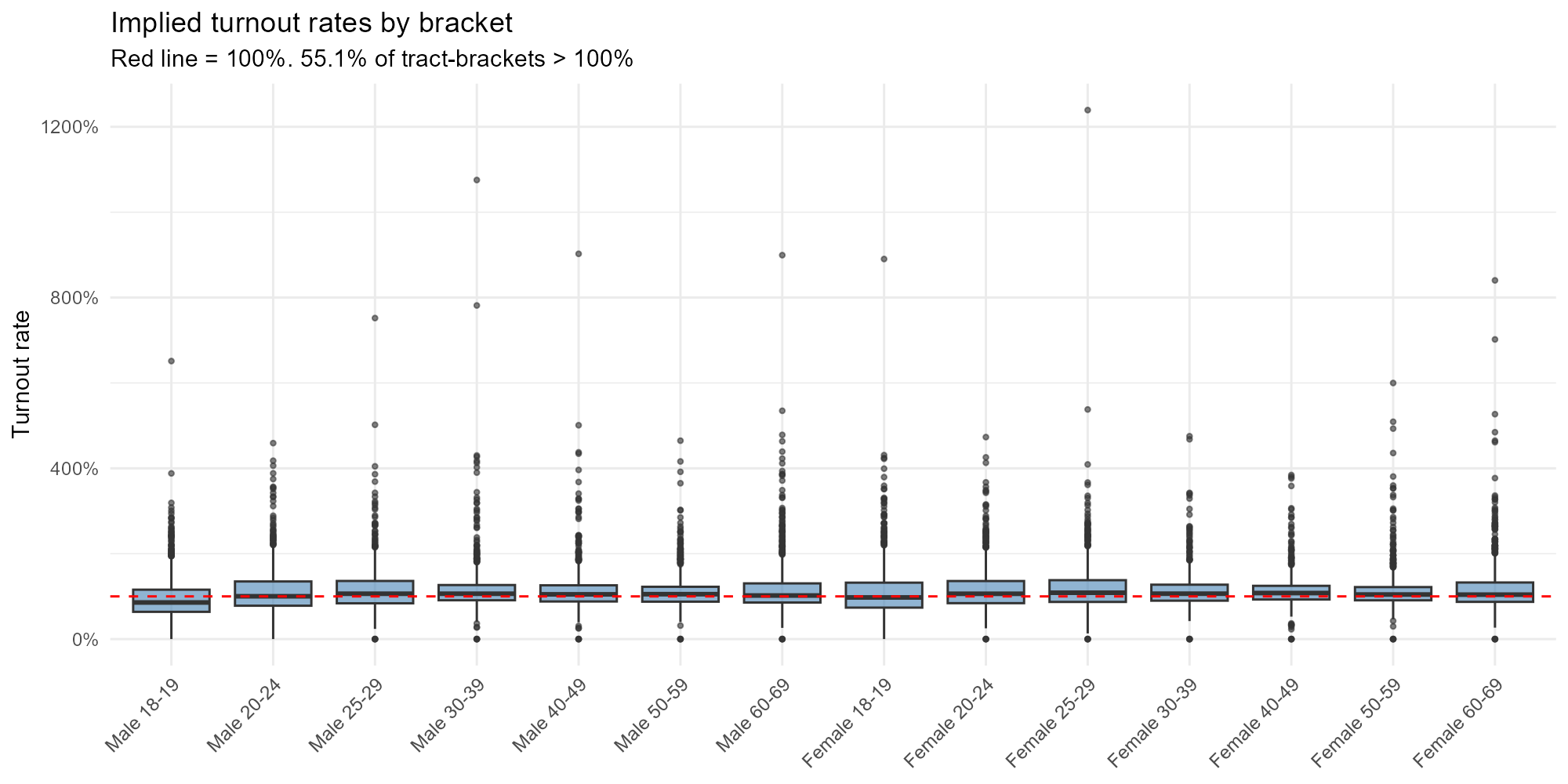
Spatial pattern:
plot_turnout_rates(result_bh, type = "map")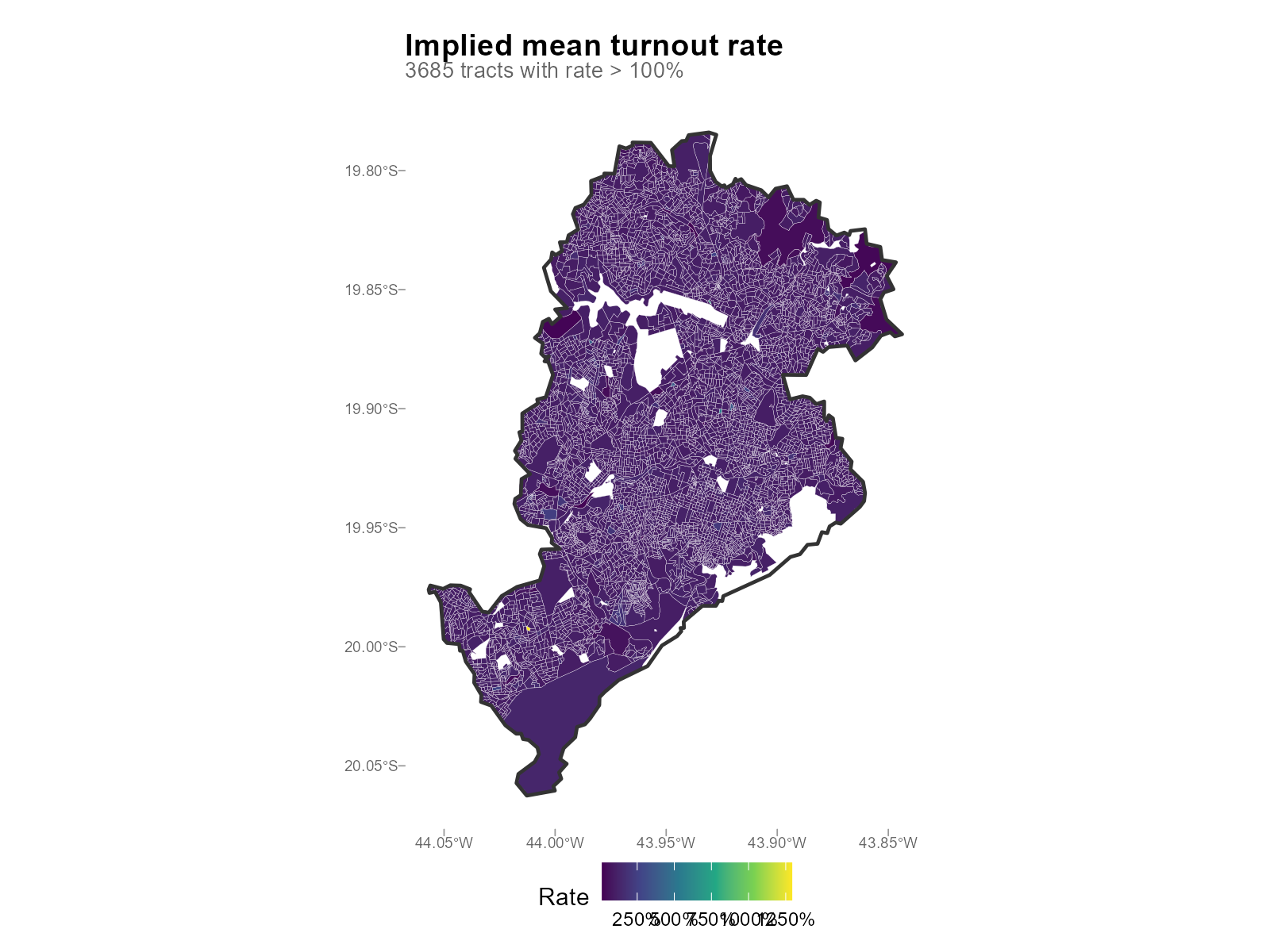
Faceted histogram:
plot_turnout_rates(result_vga, type = "histogram")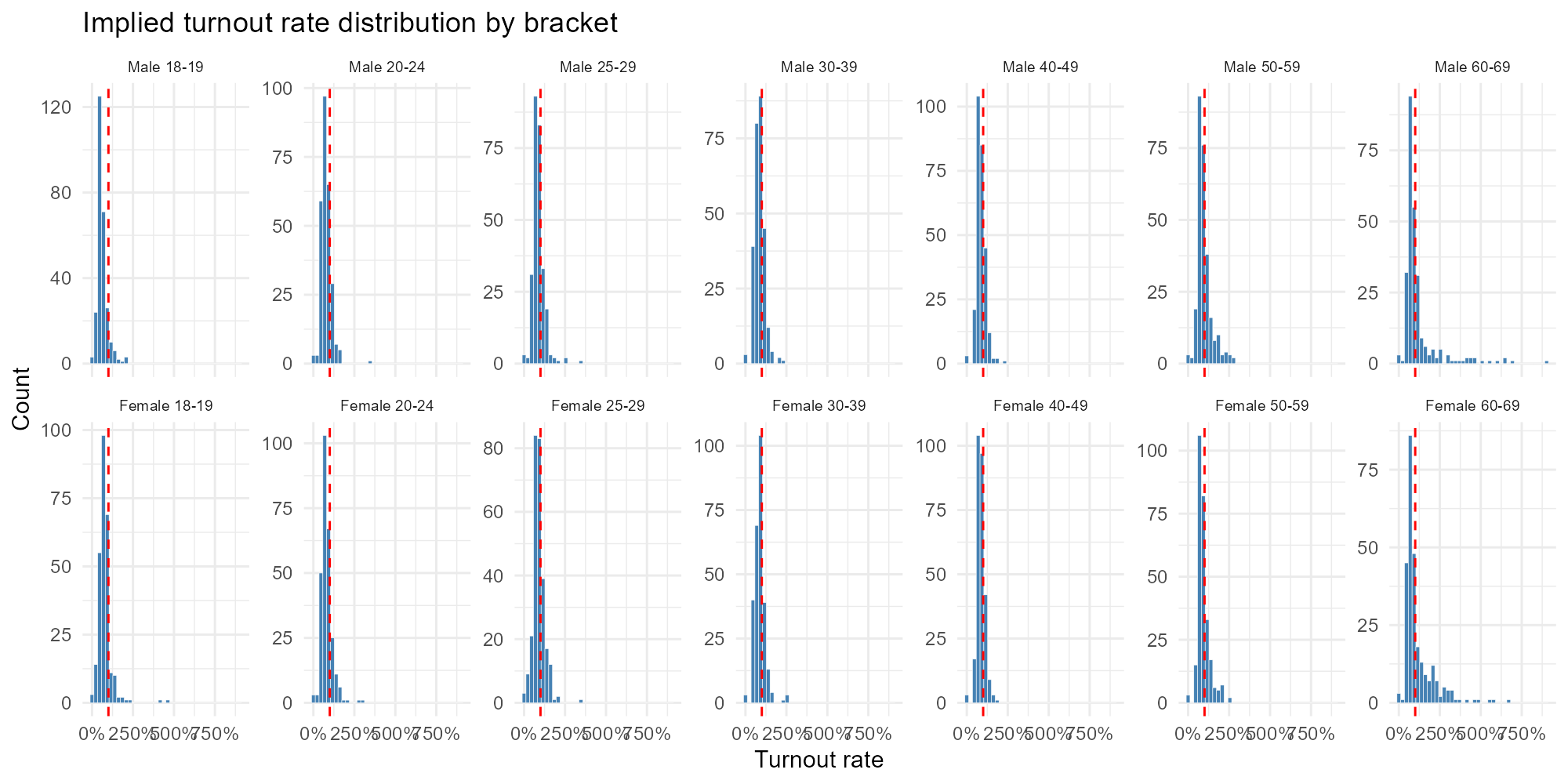
Interpretation
- Compulsory brackets clustering around 80%: Expected for ages 18–69.
- Optional brackets lower: 16–17 and 70+ should show lower turnout.
- Rates > 100%: Voters registered in one municipality who may actually reside in another (commuters, students). These boundary effects are a known limitation of the model.
8b. Income-Vote Correlation: plot_income()
Intuition
The negative correlation between PT vote share and income is one of the best-documented patterns in Brazilian political science (the “PT-income gradient”). If the interpolation preserves this gradient, it builds confidence that the spatial allocation of votes is realistic. If it destroys it, something is wrong.
Income data comes from censobr (Census Bureau microdata), which provides mean household head income by tract. No deflation is applied — within a single census year, the correlation is scale-invariant.
Applied examples
Lula (PT) vs income — expect negative correlation:
plot_income(result_vga, variable = "Lula", type = "scatter")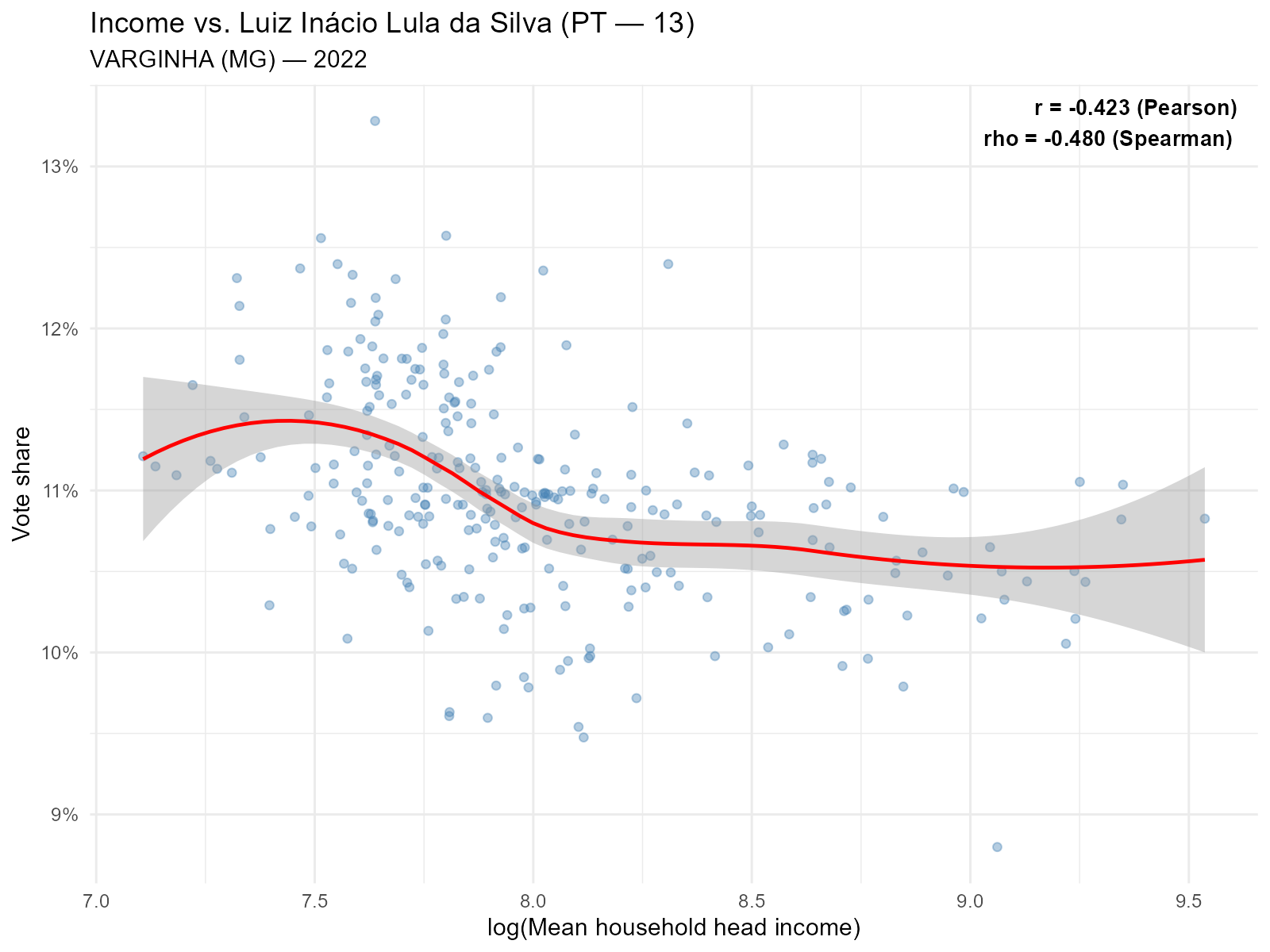
Bolsonaro (PL) vs income — expect positive correlation:
plot_income(result_vga, variable = 22, type = "scatter")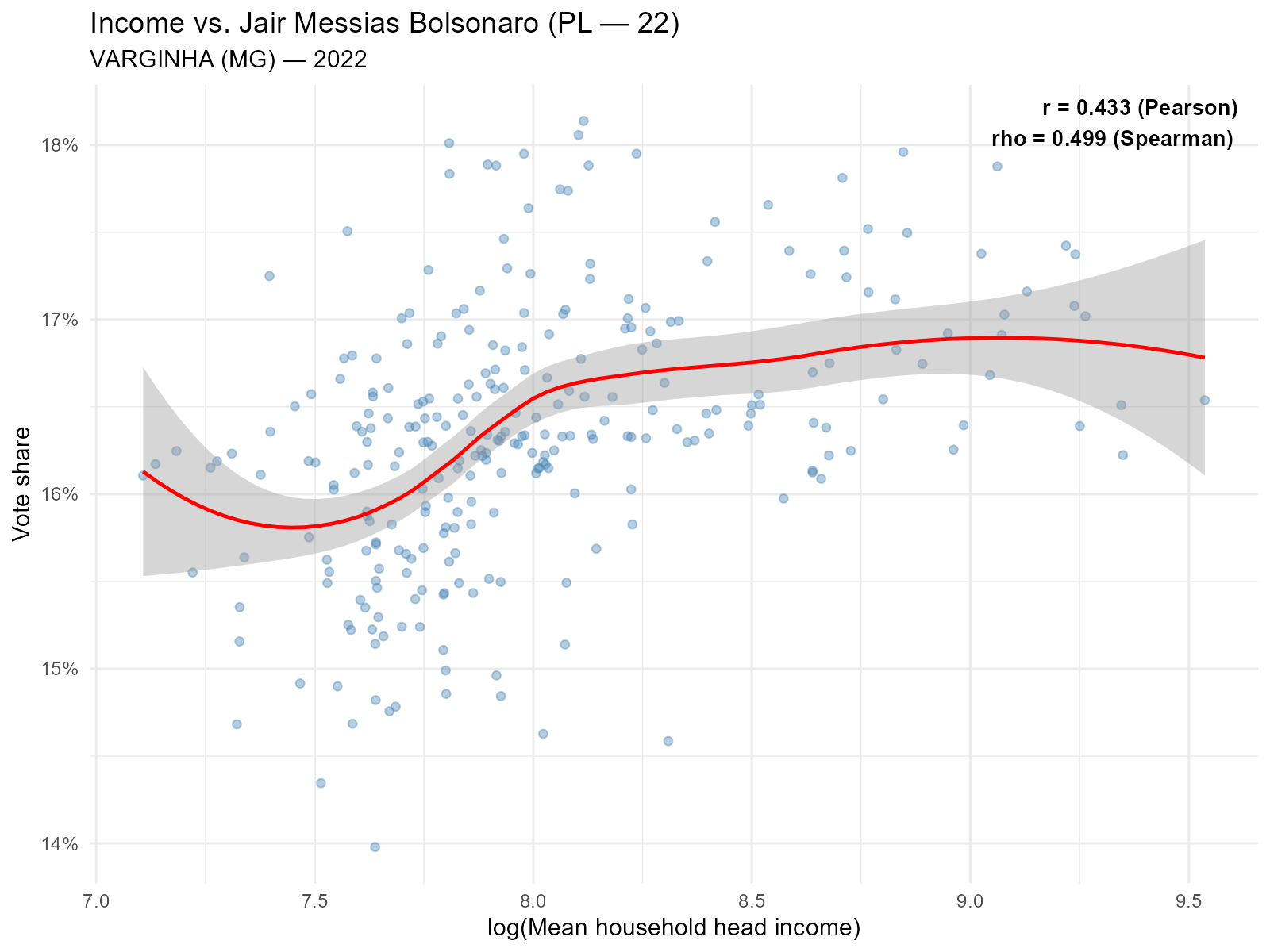
Larger city:
plot_income(result_bh, variable = "Lula", type = "scatter")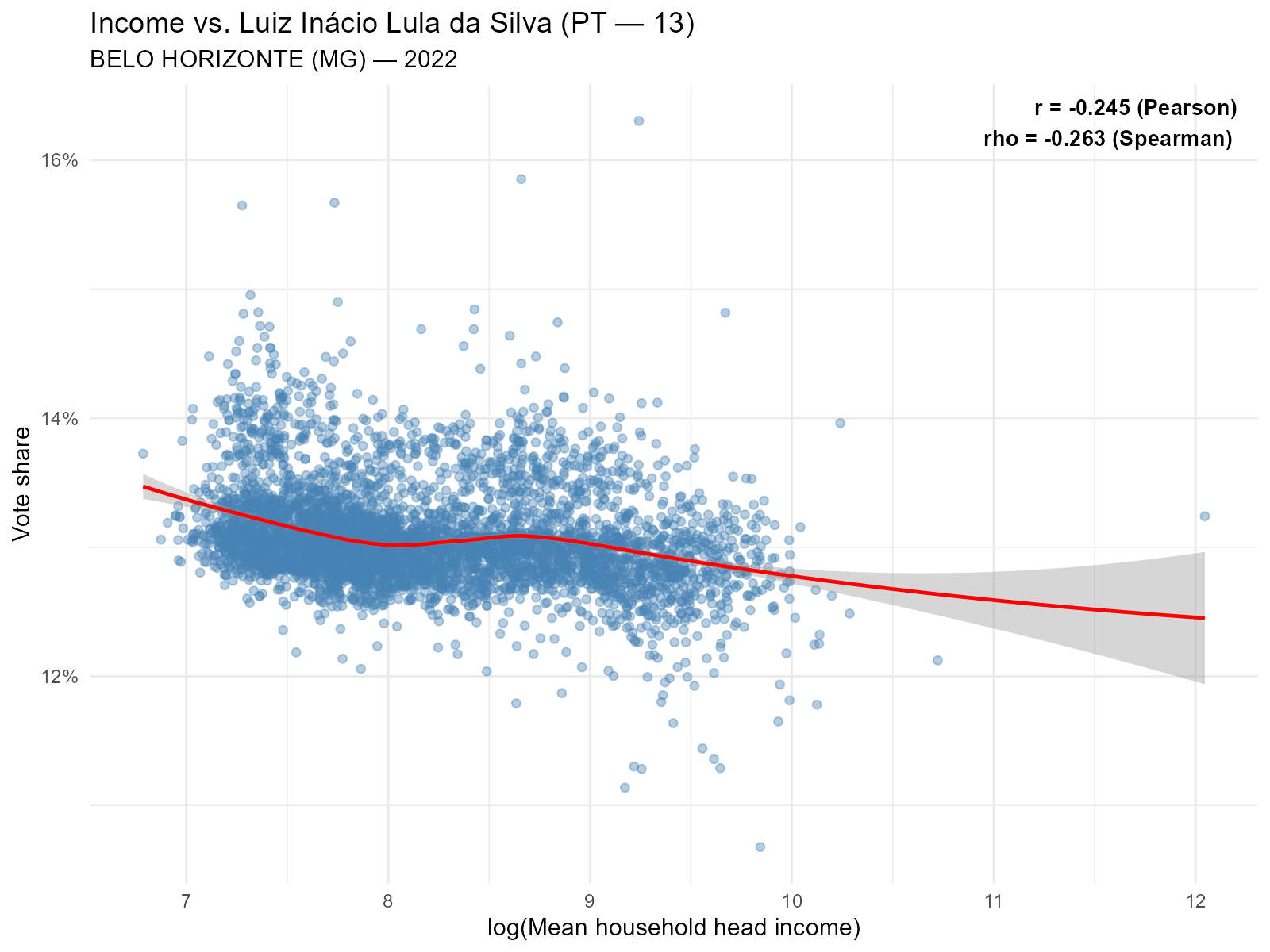
Side-by-side maps:
plot_income(result_nit, variable = "Lula", type = "map")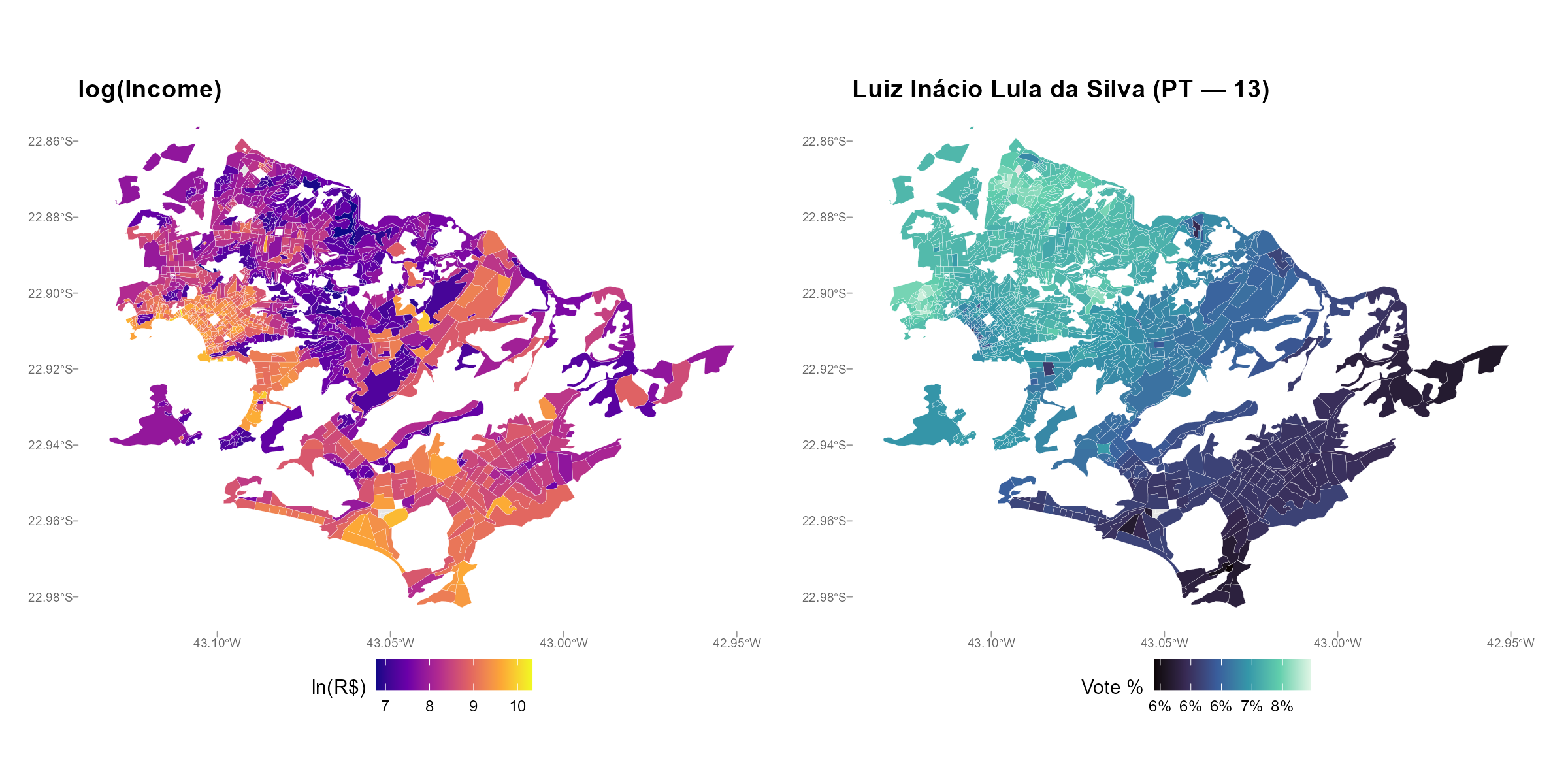
8c. Cross-Round Consistency: plot_ecological()
Intuition
Political geography is sticky. Neighborhoods that vote strongly for a candidate in round 1 should vote even more strongly in round 2 (where only 2 candidates remain). A high correlation between turno 1 and turno 2 tract-level vote shares validates that the interpolation preserves spatial patterns.
Applied examples
plot_ecological(result_vga, result_vga2,
variable = "Lula", type = "scatter")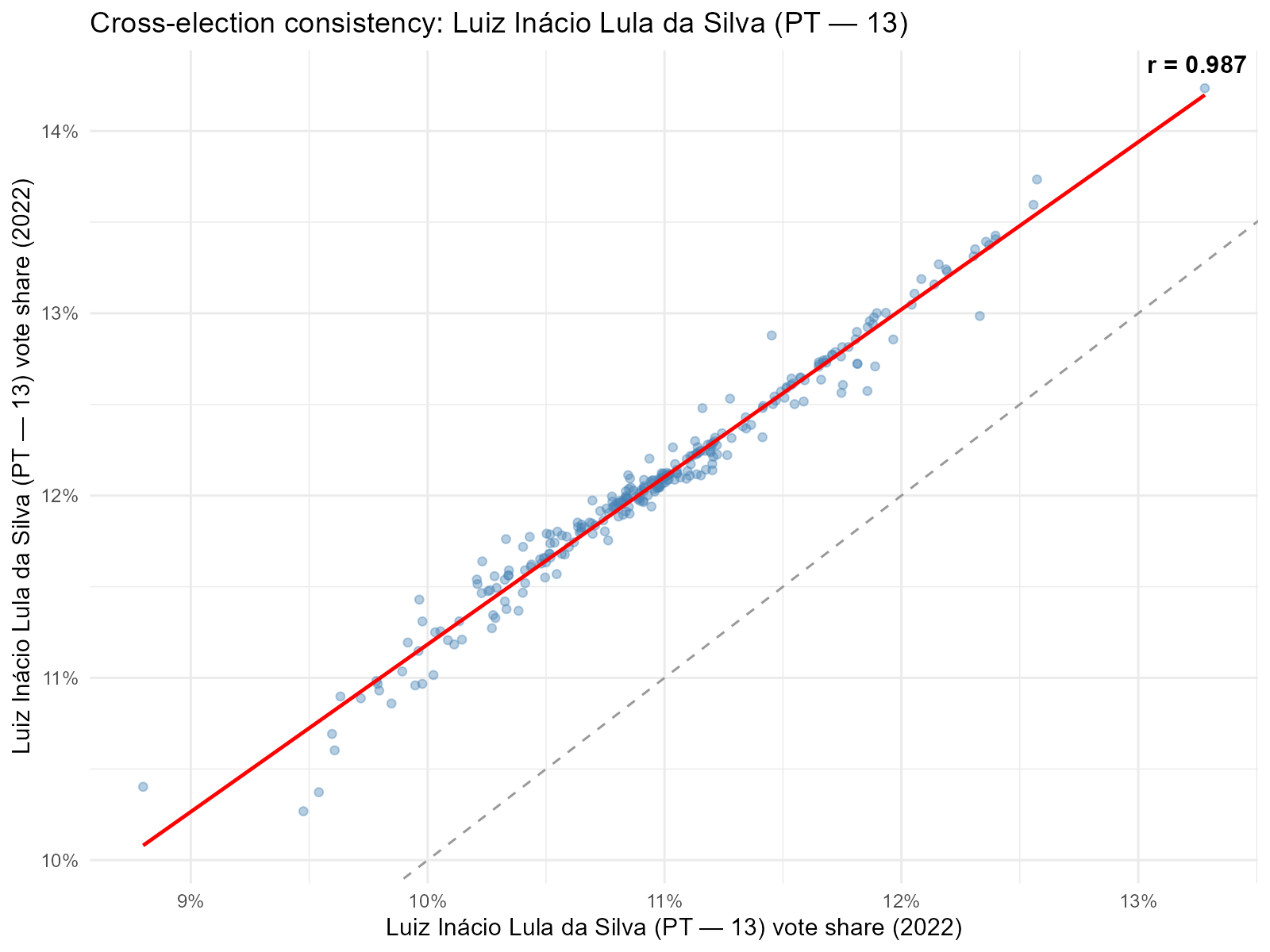
Side-by-side maps:
plot_ecological(result_vga, result_vga2,
variable = "Lula", type = "map")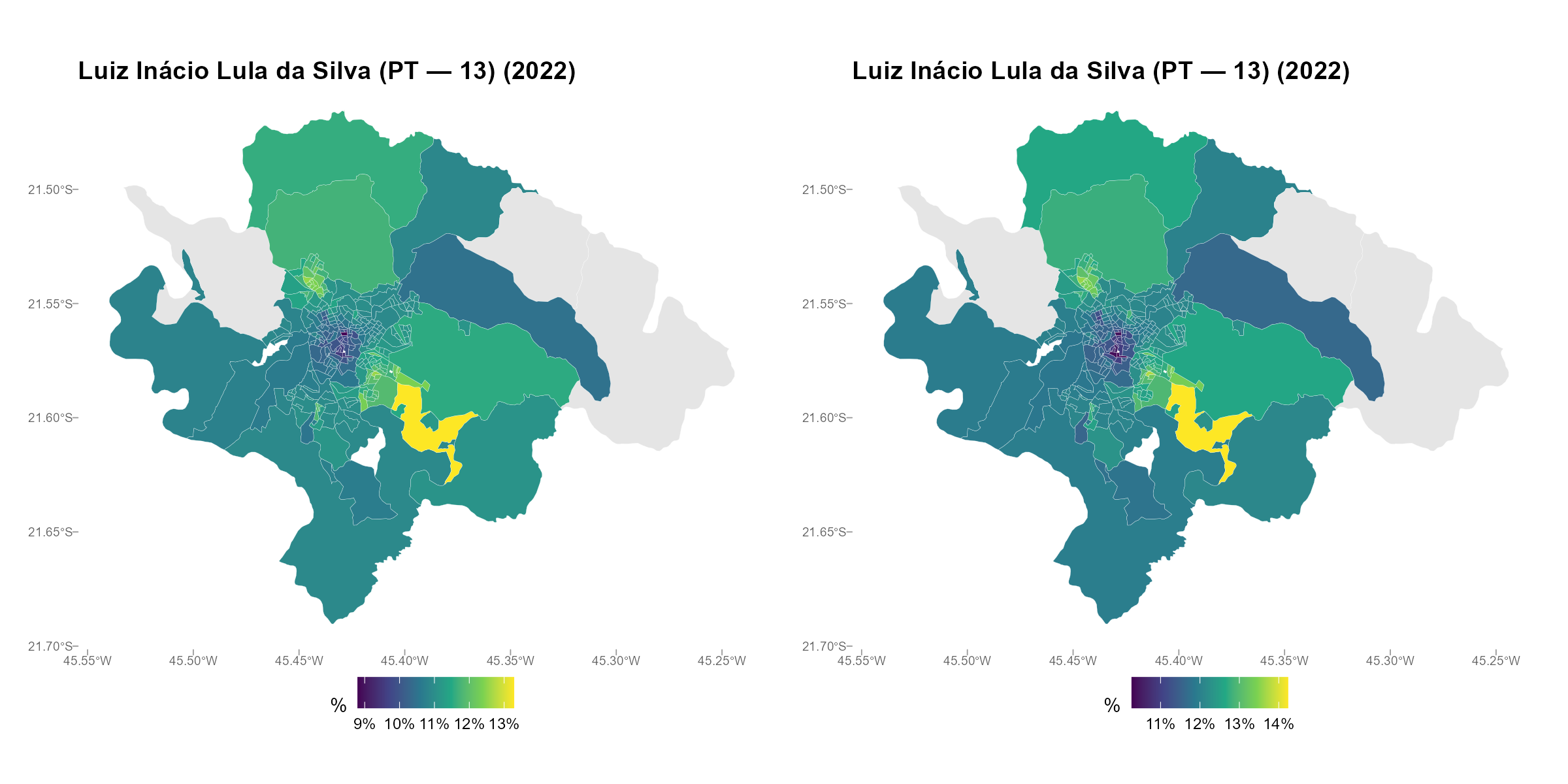
Interpretation
- Strong positive correlation between rounds: The spatial patterns are consistent, as expected. The method is recovering stable partisan geography.
- Weak or no correlation: The interpolation may be injecting noise rather than recovering signal. Investigate whether the travel time data or optimization is causing problems.
9. Quick Reference
| Function | Question it answers | Key arguments | When to worry |
|---|---|---|---|
diagnostics() |
Is anything obviously wrong? | verbose |
Any FAIL or multiple WARNs |
plot_convergence() |
Did the optimizer converge? |
which, log_y
|
Oscillating loss, non-shrinking gradient |
plot_alpha() |
How does distance decay vary? |
type, summary_fn,
brackets
|
Many tracts with |
plot_residuals() |
How well are demographics reproduced? |
type, residual_type,
summary_fn
|
Large systematic bias, Pearson > 3 |
residual_summary() |
Which brackets/tracts fit worst? | type |
One bracket much worse than others |
weight_summary() |
How concentrated are the weights? | — | Many tracts with effective N = 1 |
plot_weights() |
Where are catchment areas? |
type, tract,
top_k
|
Catchments that defy geography |
plot_connections() |
Which stations serve which tracts? |
tract, top_k,
threshold
|
Long-distance connections |
plot_travel_times() |
Is the travel time matrix clean? |
type, tract
|
Large spike at max value |
plot_residual_autocorrelation() |
Are residuals spatially clustered? |
residual_type,
summary_fn
|
Significant Moran’s I (p < 0.05) |
plot_moran() |
Is vote geography realistic? |
variable, type
|
No spatial clustering |
plot_turnout_rates() |
Are implied turnout rates plausible? | type |
Many tracts > 100% |
plot_income() |
Does the PT-income gradient hold? |
variable, type
|
Wrong sign on correlation |
plot_ecological() |
Are spatial patterns consistent? |
variable, type
|
Weak cross-round correlation |